Bone nonunion is a significant clinical problem, yet the cellular mechanisms that sustain failed repair in humans remain poorly understood. Here, we re-analyzed a human nonunion single-cell RNA sequencing (scRNA-seq) dataset to move beyond broad descriptive cell-population changes and generate a subtype-resolved, hypothesis-oriented framework. Across 16 major lineages, differential abundance testing revealed robust expansion of fibroblast, chondrocyte, and macrophage neighborhoods in nonunion. Intercellular communication reconstruction further showed that macrophages were the dominant upregulated signal “senders”, whereas fibroblasts were the primary “receivers”, with a disease-enhanced macrophage-to-fibroblast TGFB1–(TGFBR1+TGFBR2) axis forming a central hub. Fibroblasts in nonunion also showed metabolic rewiring and an increased senescence signature, including strengthened metabolite-mediated macrophage-to-fibroblast signaling (e.g., glutamine-linked pathways). Further subclustering identified a disease-enriched fibroblast “Fib1 executor” state that acts as a pro-fibrotic signaling sink, governed by a 13-transcription-factor regulon and a compact five-gene effector module (ACTA2, F3, THBS1, SERPINE1, NTM). Finally, we performed an exploratory computational compound-prioritization analysis linked to the Fib1-centered program, generating preliminary hypotheses for future validation rather than validated therapeutic nominations. Together, our results suggest that human nonunion contains a disease-enriched macrophage-to-fibroblast communication program and a fibroblast-centered effector signature that may help guide future mechanistic and therapeutic studies.
- Open Access
- Article
Single-Cell Re-Mining Identifies a Pathogenic Fibroblast Subpopulation and Nominates Exploratory Therapeutic Hypotheses in Human Bone Nonunion
Author Information
Received: 23 Feb 2026 | Revised: 06 Apr 2026 | Accepted: 14 Apr 2026 | Published: 27 Apr 2026
Abstract
Graphical Abstract
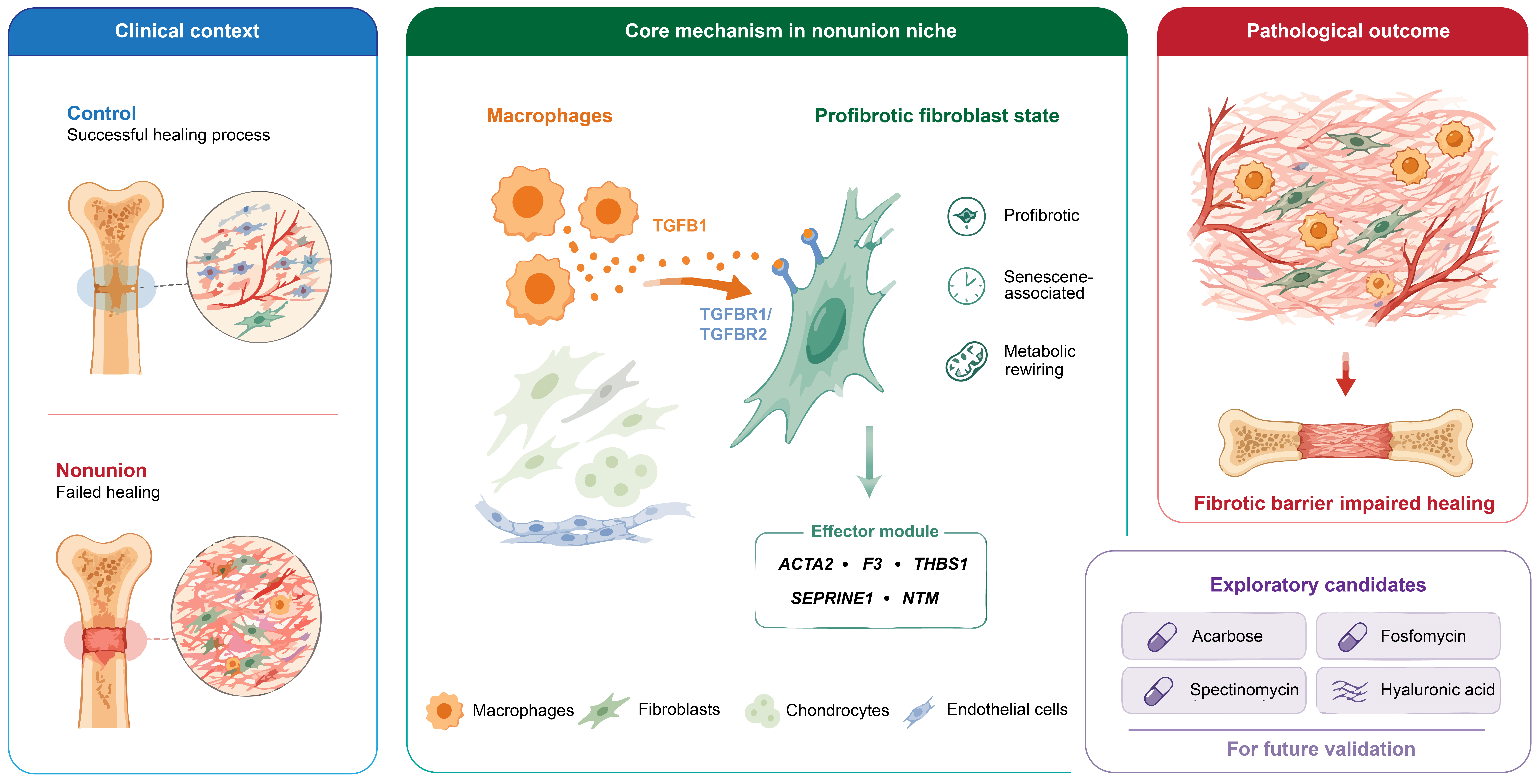
Keywords
bone nonunion | single-cell RNA sequencing | fibroblast | drug repurposing
References
- 1.
Vanderkarr, M.F.; Ruppenkamp, J.W.; Vanderkarr, M.; et al. Risk factors and healthcare costs associated with long bone fracture non-union: A retrospective US claims database analysis. J. Orthop. Surg. Res. 2023, 18, 745. https://doi.org/10.1186/s13018-023-04232-3.
- 2.
Thomas, J.D.; Kehoe, J.L. Bone Nonunion. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2025.
- 3.
Vincken, L.; van der Broeck, L.; Geurts, J.; et al. The effect of post-traumatic long bone non-unions on health-related quality of life. Injury 2023, 54, 110929. https://doi.org/10.1016/j.injury.2023.110929.
- 4.
Saul, D.; Menger, M.M.; Ehnert, S.; et al. Bone Healing Gone Wrong: Pathological Fracture Healing and Non-Unions-Overview of Basic and Clinical Aspects and Systematic Review of Risk Factors. Bioengineering 2023, 10, 85. https://doi.org/10.3390/bioengineering10010085.
- 5.
Capobianco, C.A.; Hankenson, K.D.; Knights, A.J. Temporal dynamics of immune-stromal cell interactions in fracture healing. Front. Immunol. 2024, 15, 1352819. https://doi.org/10.3389/fimmu.2024.1352819.
- 6.
Wildemann, B.; Ignatius, A.; Leung, F.; et al. Non-union bone fractures. Nat. Rev. Dis. Primers 2021, 7, 57. https://doi.org/10.1038/s41572-021-00289-8.
- 7.
Vukicevic, S.; Peric, M.; Bordukalo-Niksic, T.; et al. Time to consider fracture nonunion an orphan disease? An update into pathophysiology, epidemiology and therapeutic solutions. Eur. J. Trauma Emerg. Surg. 2025, 51, 255. https://doi.org/10.1007/s00068-025-02918-3.
- 8.
Qin, Q.; Lee, S.; Patel, N.; et al. Neurovascular coupling in bone regeneration. Exp. Mol. Med. 2022, 54, 1844–1849. https://doi.org/10.1038/s12276-022-00899-6.
- 9.
Panteli, M.; Pountos, I.; Jones, E.; et al. Biological and molecular profile of fracture non-union tissue: Current insights. J. Cell. Mol. Med. 2015, 19, 685–713. https://doi.org/10.1111/jcmm.12532.
- 10.
Avin, K.G.; Dominguez, J.M., 2nd; Chen, N.X.; et al. Single-cell RNAseq provides insight into altered immune cell populations in human fracture nonunions. J. Orthop. Res. 2023, 41, 1060–1069. https://doi.org/10.1002/jor.25452.
- 11.
La Manna, F.; Hanhart, D.; Kloen, P.; et al. Molecular profiling of osteoprogenitor cells reveals FOS as a master regulator of bone non-union. Gene 2023, 874, 147481. https://doi.org/10.1016/j.gene.2023.147481.
- 12.
Salichos, L.; Thayavally, R.; Kloen, P.; et al. Human nonunion tissues display differential gene expression in comparison to physiological fracture callus. Bone 2024, 183, 117091. https://doi.org/10.1016/j.bone.2024.117091.
- 13.
Hao, Y.; Stuart, T.; Kowalski, M.H.; et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 2024, 42, 293–304. https://doi.org/10.1038/s41587-023-01767-y.
- 14.
Zeng, Z.; Ma, Y.; Hu, L.; et al. OmicVerse: A framework for bridging and deepening insights across bulk and single-cell sequencing. Nat. Commun. 2024, 15, 5983. https://doi.org/10.1038/s41467-024-50194-3.
- 15.
Germain, P.L.; Lun, A.; Garcia Meixide, C.; et al. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res 2021, 10, 979. https://doi.org/10.12688/f1000research.73600.2.
- 16.
Korsunsky, I.; Millard, N.; Fan, J.; et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 2019, 16, 1289–1296. https://doi.org/10.1038/s41592-019-0619-0.
- 17.
Dann, E.; Henderson, N.C.; Teichmann, S.A.; et al. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 2022, 40, 245–253. https://doi.org/10.1038/s41587-021-01033-z.
- 18.
Chu, T.; Wang, Z.; Pe’er, D.; et al. Cell type and gene expression deconvolution with BayesPrism enables Bayesian integrative analysis across bulk and single-cell RNA sequencing in oncology. Nat. Cancer 2022, 3, 505–517. https://doi.org/10.1038/s43018-022-00356-3.
- 19.
Jin, S.; Plikus, M.V.; Nie, Q. CellChat for systematic analysis of cell–cell communication from single-cell transcriptomics. Nat. Protoc. 2025, 20, 180–219. https://doi.org/10.1038/s41596-024-01045-4.
- 20.
Browaeys, R.; Saelens, W.; Saeys, Y. NicheNet: Modeling intercellular communication by linking ligands to target genes. Nat. Methods 2020, 17, 159–162. https://doi.org/10.1038/s41592-019-0667-5.
- 21.
Alghamdi, N.; Chang, W.; Dang, P.; et al. A graph neural network model to estimate cell-wise metabolic flux using single-cell RNA-seq data. Genome Res. 2021, 31, 1867–1884. https://doi.org/10.1101/gr.271205.120.
- 22.
Zheng, R.; Zhang, Y.; Tsuji, T.; et al. MEBOCOST maps metabolite-mediated intercellular communications using single-cell RNA-seq. Nucleic Acids Res. 2025, 53, gkaf569. https://doi.org/10.1093/nar/gkaf569.
- 23.
Wang, J.; Zhou, X.; Yu, P.; et al. A transcriptome-based human universal senescence index (hUSI) robustly predicts cellular senescence under various conditions. Nat. Aging 2025, 5, 1159–1175. https://doi.org/10.1038/s43587-025-00886-2.
- 24.
Bauer-Rowe, K.E.; Pham, B.; Griffin, M.; et al. Creeping fat-derived mechanosensitive fibroblasts drive intestinal fibrosis in Crohn’s disease strictures. Cell 2025, 188, 6536–6553.e26. https://doi.org/10.1016/j.cell.2025.08.029.
- 25.
Aibar, S.; González-Blas, C.B.; Moerman, T.; et al. SCENIC: Single-cell regulatory network inference and clustering. Nat. Methods 2017, 14, 1083–1086. https://doi.org/10.1038/nmeth.4463.
- 26.
Zhou, Y.; Zhou, B.; Pache, L.; et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1523. https://doi.org/10.1038/s41467-019-09234-6.
- 27.
Osorio, D.; Zhong, Y.; Li, G.; et al. scTenifoldKnk: An efficient virtual knockout tool for gene function predictions via single-cell gene regulatory network perturbation. Patterns 2022, 3, 100434. https://doi.org/10.1016/j.patter.2022.100434.
- 28.
Liu, L.; Li, W.; Wang, F.; et al. A pre-trained large generative model for translating single-cell transcriptomes to proteomes. Nat. Biomed. Eng. 2025. https://doi.org/10.1038/s41551-025-01528-z.
- 29.
Knox, C.; Wilson, M.; Klinger, C.M.; et al. DrugBank 6.0: The DrugBank Knowledgebase for 2024. Nucleic Acids Res. 2024, 52, D1265–D1275. https://doi.org/10.1093/nar/gkad976.
- 30.
UniProt Consortium. UniProt: The Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 2025, 53, D609–D617. https://doi.org/10.1093/nar/gkae1010.
- 31.
Zdrazil, B.; Felix, E.; Hunter, F.; et al. The ChEMBL Database in 2023: A drug discovery platform spanning multiple bioactivity data types and time periods. Nucleic Acids Res. 2024, 52, D1180–D1192. https://doi.org/10.1093/nar/gkad1004.
- 32.
He, H.; Chen, G.; Chen, C.Y. NHGNN-DTA: A node-adaptive hybrid graph neural network for interpretable drug-target binding affinity prediction. Bioinformatics 2023, 39, btad355. https://doi.org/10.1093/bioinformatics/btad355.
- 33.
Passaro, S.; Corso, G.; Wohlwend, J.; et al. Boltz-2: Towards Accurate and Efficient Binding Affinity Prediction. bioRxiv 2025. https://doi.org/10.1101/2025.06.14.659707.
- 34.
Li, Z.; Jiang, J.; Cai, K.; et al. CCN2 mediates fibroblast-macrophage interaction in knee arthrofibrosis based on single-cell RNA-seq analysis. Bone Res. 2025, 13, 26. https://doi.org/10.1038/s41413-025-00400-9.
- 35.
Watanabe, N.; Yoshida, M.; Hirano, Y.; et al. Integrated spatial and single-cell transcriptomics reveal PAK kinase as a therapeutic target in fibroblastic foci and dense fibrosis of idiopathic pulmonary fibrosis. Eur. Respir. J. 2025, 66, 2400022. https://doi.org/10.1183/13993003.00022-2025.
- 36.
Chen, H.; You, R.; Guo, J.; et al. WWP2 Regulates Renal Fibrosis and the Metabolic Reprogramming of Profibrotic Myofibroblasts. J. Am. Soc. Nephrol. 2024, 35, 696–718. https://doi.org/10.1681/asn.0000000000000328.
- 37.
Ge, C.; Huang, M.; Han, Y.; et al. Demethyleneberberine Alleviates Pulmonary Fibrosis through Disruption of USP11 Deubiquitinating GREM1. Pharmaceuticals 2024, 17, 279. https://doi.org/10.3390/ph17030279.
- 38.
Luzina, I.G.; Rus, V.; Lockatell, V.; et al. Regulator of Cell Cycle Protein (RGCC/RGC-32) Protects against Pulmonary Fibrosis. Am. J. Respir. Cell Mol. Biol. 2022, 66, 146–157. https://doi.org/10.1165/rcmb.2021-0022OC.
- 39.
Zhou, J.; An, X.; Xia, X.; et al. Aging-associated interleukin-11 drives the molecular mechanism and targeted therapy of idiopathic pulmonary fibrosis. Eur. J. Med. Res. 2025, 30, 542. https://doi.org/10.1186/s40001-025-02755-5.
- 40.
Kuppe, C.; Ibrahim, M.M.; Kranz, J.; et al. Decoding myofibroblast origins in human kidney fibrosis. Nature 2021, 589, 281–286. https://doi.org/10.1038/s41586-020-2941-1.
- 41.
Li, L.; Tao, M.; Gao, X.; et al. Uncovering key markers and therapeutic targets for renal fibrosis in diabetic kidney disease through bulk and single-cell RNA sequencing. J. Transl. Med. 2025, 23, 742. https://doi.org/10.1186/s12967-025-06554-8.
- 42.
Pokatayev, V.; Jaiswal, A.; Shih, A.R.; et al. Bidirectional CRISPR screens decode a GLIS3-dependent fibrotic cell circuit. Nature 2026, 650, 997–1006. https://doi.org/10.1038/s41586-025-09907-x.
- 43.
Liu, J.; Zhang, J.; Lin, X.; et al. Age-associated callus senescent cells produce TGF-β1 that inhibits fracture healing in aged mice. J. Clin. Investig. 2022, 132, e148073. https://doi.org/10.1172/jci148073.
- 44.
Wu, M.; Wu, S.; Chen, W.; et al. The roles and regulatory mechanisms of TGF-β and BMP signaling in bone and cartilage development, homeostasis and disease. Cell Res. 2024, 34, 101–123. https://doi.org/10.1038/s41422-023-00918-9.
- 45.
Khalil, N.; Parekh, T.V.; O’Connor, R.; et al. Regulation of the effects of TGF-β1 by activation of latent TGF-β1 and differential expression of TGF-β receptors (TβR-I and TβR-II) in idiopathic pulmonary fibrosis. Thorax 2001, 56, 907–915. https://doi.org/10.1136/thorax.56.12.907.
- 46.
Zheng, N.; Xu, J.; Ruan, Y.C.; et al. Magnesium facilitates the healing of atypical femoral fractures: A single-cell transcriptomic study. Mater. Today 2022, 52, 43–62. https://doi.org/10.1016/j.mattod.2021.11.028.
- 47.
Bowers, K.M.; Anderson, D.E. Delayed Union and Nonunion: Current Concepts, Prevention, and Correction: A Review. Bioengineering 2024, 11, 525. https://doi.org/10.3390/bioengineering11060525.
- 48.
Julien, A.; Kanagalingam, A.; Martínez-Sarrà, E.; et al. Direct contribution of skeletal muscle mesenchymal progenitors to bone repair. Nat. Commun. 2021, 12, 2860. https://doi.org/10.1038/s41467-021-22842-5.
- 49.
He, Q.; Lu, J.; Liang, Q.; et al. Prg4+ fibroadipogenic progenitors in muscle are crucial for bone fracture repair. Proc. Natl. Acad. Sci. U.S.A. 2025, 122, e2417806122. https://doi.org/10.1073/pnas.2417806122.
- 50.
Dupont, S.; Morsut, L.; Aragona, M.; et al. Role of YAP/TAZ in mechanotransduction. Nature 2011, 474, 179–183. https://doi.org/10.1038/nature10137.
- 51.
Tai, Y.; Woods, E.L.; Dally, J.; et al. Myofibroblasts: Function, Formation, and Scope of Molecular Therapies for Skin Fibrosis. Biomolecules 2021, 11, 1095. https://doi.org/10.3390/biom11081095.
- 52.
Huang, F.; Wei, G.; Wang, H.; et al. Fibroblasts inhibit osteogenesis by regulating nuclear-cytoplasmic shuttling of YAP in mesenchymal stem cells and secreting DKK1. Biol. Res. 2024, 57, 4. https://doi.org/10.1186/s40659-023-00481-y.
- 53.
Wang, L.; Tower, R.J.; Chandra, A.; et al. Periosteal Mesenchymal Progenitor Dysfunction and Extraskeletally-Derived Fibrosis Contribute to Atrophic Fracture Nonunion. J. Bone Miner. Res. 2019, 34, 520–532. https://doi.org/10.1002/jbmr.3626.
- 54.
Saul, D.; Monroe, D.G.; Rowsey, J.L.; et al. Modulation of fracture healing by the transient accumulation of senescent cells. eLife 2021, 10, e69958. https://doi.org/10.7554/eLife.69958.
- 55.
Trogisch, F.A.; Abouissa, A.; Keles, M.; et al. Endothelial cells drive organ fibrosis in mice by inducing expression of the transcription factor SOX9. Sci. Transl. Med. 2024, 16, eabq4581. https://doi.org/10.1126/scitranslmed.abq4581.
- 56.
Bai, W.W.; Tang, Z.Y.; Shan, T.C.; et al. Up-regulation of paired-related homeobox 2 promotes cardiac fibrosis in mice following myocardial infarction by targeting of Wnt5a. J. Cell. Mol. Med. 2020, 24, 2319–2329. https://doi.org/10.1111/jcmm.14914.
- 57.
Henley, M.J.; Koehler, A.N. Advances in targeting ‘undruggable’ transcription factors with small molecules. Nat. Rev. Drug Discov. 2021, 20, 669–688. https://doi.org/10.1038/s41573-021-00199-0.
- 58.
Joris, V.; Schumacher, A.; Marks, M.P.; et al. FGF9 treatment reduces off-target chondrocytes from iPSC-derived kidney organoids. npj Regen. Med. 2025, 10, 41. https://doi.org/10.1038/s41536-025-00428-9.
- 59.
Blanco-Mezquita, J.T.; Hutcheon, A.E.; Zieske, J.D. Role of thrombospondin-1 in repair of penetrating corneal wounds. Investig. Ophthalmol. Vis. Sci. 2013, 54, 6262–6268. https://doi.org/10.1167/iovs.13-11710.
- 60.
Shinde, A.V.; Humeres, C.; Frangogiannis, N.G. The role of α-smooth muscle actin in fibroblast-mediated matrix contraction and remodeling. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 298–309. https://doi.org/10.1016/j.bbadis.2016.11.006.
- 61.
Simone, T.M.; Higgins, C.E.; Czekay, R.P.; et al. SERPINE1: A Molecular Switch in the Proliferation-Migration Dichotomy in Wound-“Activated” Keratinocytes. Adv. Wound Care 2014, 3, 281–290. https://doi.org/10.1089/wound.2013.0512.
- 62.
Webber, J.; Meran, S.; Steadman, R.; et al. Hyaluronan orchestrates transforming growth factor-β1-dependent maintenance of myofibroblast phenotype. J. Biol. Chem. 2009, 284, 9083–9092. https://doi.org/10.1074/jbc.M806989200.
- 63.
Wang, B.; Han, J.; Elisseeff, J.H.; et al. The senescence-associated secretory phenotype and its physiological and pathological implications. Nat. Rev. Mol. Cell Biol. 2024, 25, 958–978. https://doi.org/10.1038/s41580-024-00727-x.
- 64.
Feng, L.; Chen, X.; Huang, Y.; et al. Immunometabolism changes in fibrosis: From mechanisms to therapeutic strategies. Front. Pharmacol. 2023, 14, 1243675. https://doi.org/10.3389/fphar.2023.1243675.
- 65.
Farah, H.; Young, S.P.; Mauro, C.; et al. Metabolic dysfunction and inflammatory disease: The role of stromal fibroblasts. FEBS J. 2021, 288, 5555–5568. https://doi.org/10.1111/febs.15644.
- 66.
Schlundt, C.; El Khassawna, T.; Serra, A.; et al. Macrophages in bone fracture healing: Their essential role in endochondral ossification. Bone 2018, 106, 78–89. https://doi.org/10.1016/j.bone.2015.10.019.
- 67.
Cherief, M.; Xu, J.; Li, Z.; et al. TrkA-mediated sensory innervation of injured mouse tendon supports tendon sheath progenitor cell expansion and tendon repair. Sci. Transl. Med. 2023, 15, eade4619. https://doi.org/10.1126/scitranslmed.ade4619.
- 68.
Laurencin, C.T.; Ashe, K.M.; Henry, N.; et al. Delivery of small molecules for bone regenerative engineering: Preclinical studies and potential clinical applications. Drug Discov. Today 2014, 19, 794–800. https://doi.org/10.1016/j.drudis.2014.01.012.
- 69.
Li, X.; Zheng, S.; Xu, H.; et al. The direct and indirect inhibition of proinflammatory adipose tissue macrophages by acarbose in diet-induced obesity. Cell Rep. Med. 2025, 6, 101883. https://doi.org/10.1016/j.xcrm.2024.101883.
- 70.
Hung, T.W.; Yu, M.H.; Yang, T.Y.; et al. Acarbose Protects Glucolipotoxicity-Induced Diabetic Nephropathy by Inhibiting Ras Expression in High-Fat Diet-Fed db/db Mice. Int. J. Mol. Sci. 2022, 23, 15312. https://doi.org/10.3390/ijms232315312.
- 71.
Zeitlinger, M.; Marsik, C.; Steiner, I.; et al. Immunomodulatory effects of fosfomycin in an endotoxin model in human blood. J. Antimicrob. Chemother. 2007, 59, 219–223. https://doi.org/10.1093/jac/dkl464.
- 72.
Schintler, M.V.; Traunmüller, F.; Metzler, J.; et al. High fosfomycin concentrations in bone and peripheral soft tissue in diabetic patients presenting with bacterial foot infection. J. Antimicrob. Chemother. 2009, 64, 574–578. https://doi.org/10.1093/jac/dkp230.
- 73.
Lam, T.; Brennan, M.D.; Morrison, D.A.; et al. Femtoliter droplet confinement of Streptococcus pneumoniae: Bacterial genetic transformation by cell-cell interaction in droplets. Lab Chip 2019, 19, 682–692. https://doi.org/10.1039/c8lc01367e.
- 74.
Wang, X.; Balaji, S.; Steen, E.H.; et al. High-molecular weight hyaluronan attenuates tubulointerstitial scarring in kidney injury. JCI Insight 2020, 5, e136345. https://doi.org/10.1172/jci.insight.136345.
- 75.
Schumann, C. Medical, nutritional and technological properties of lactulose. An update. Eur. J. Nutr. 2002, 41, I17–I25. https://doi.org/10.1007/s00394-002-1103-6.
- 76.
Ip, W.K.; Medzhitov, R. Macrophages monitor tissue osmolarity and induce inflammatory response through NLRP3 and NLRC4 inflammasome activation. Nat. Commun. 2015, 6, 6931. https://doi.org/10.1038/ncomms7931.

This work is licensed under a Creative Commons Attribution 4.0 International License.


