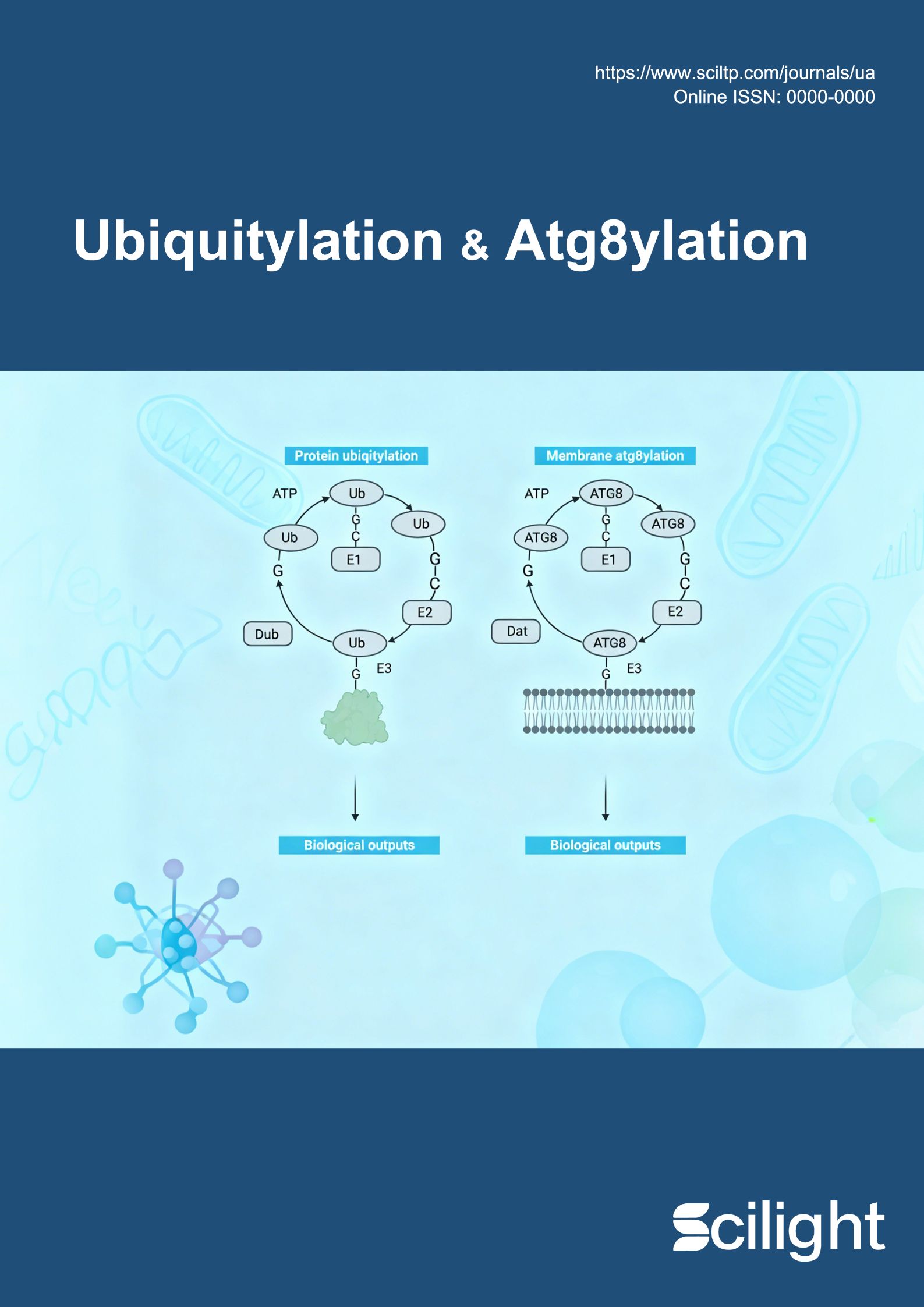
Protein ATG8ylation is a post-translational modification (PTM) wherein ubiquitin-like ATG8 proteins covalently conjugate to cellular proteins. In our recent work, we identified ATG7 as a principal substrate of this modification, with K140 as the major site, and demonstrated that ATG8ylation of ATG7 serves as an endogenous regulatory brake on autophagy by disrupting its interaction with ATG3. In this perspective article, we briefly overview the evolution of the protein ATG8ylation field—from its initial discovery to the mechanistic dissection of its core machinery—with emphasis on our recent functional characterization of ATG7 as both enzyme and substrate. We then discuss key unanswered questions, including the search for putative E3 ligases, the stress-responsive landscape of protein ATG8ylation, and emerging links to human disease.
- Open Access
- Perspective
Feedback at the Core: Protein ATG8ylation as a Regulatory Brake on the Autophagy Engine
- Huazhong Xie,
- Min Li *
Author Information
Received: 11 Feb 2026 | Revised: 07 Mar 2026 | Accepted: 11 Mar 2026 | Published: 01 Apr 2026
Abstract
Keywords
References
- 1.
Choi, A.M.; Ryter, S.W.; Levine, B. Autophagy in human health and disease. N. Engl. J. Med. 2013, 368, 651–662. https://doi.org/10.1056/NEJMra1205406.
- 2.
Levine, B.; Kroemer, G. Biological Functions of Autophagy Genes: A Disease Perspective. Cell 2019, 176, 11–42. https://doi.org/10.1016/j.cell.2018.09.048.
- 3.
Klionsky, D.J.; Schulman, B.A. Dynamic regulation of macroautophagy by distinctive ubiquitin-like proteins. Nat. Struct. Mol. Biol. 2014, 21, 336–345. https://doi.org/10.1038/nsmb.2787.
- 4.
Kumar, S.; Jia, J.; Deretic, V. Atg8ylation as a general membrane stress and remodeling response. Cell Stress 2021, 5, 128–142. https://doi.org/10.15698/cst2021.09.255.
- 5.
Deretic, V.; Lazarou, M. A guide to membrane Atg8ylation and autophagy with reflections on immunity. J. Cell Biol. 2022, 221, e202203083. https://doi.org/10.1083/jcb.202203083.
- 6.
Ichimura, Y.; Kirisako, T.; Takao, T.; et al. A ubiquitin-like system mediates protein lipidation. Nature 2000, 408, 488–492. https://doi.org/10.1038/35044114.
- 7.
Fujita, N.; Itoh, T.; Omori, H.; et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell 2008, 19, 2092–2100. https://doi.org/10.1091/mbc.e07-12-1257.
- 8.
Taherbhoy, A.M.; Tait, S.W.; Kaiser, S.E.; et al. Atg8 transfer from Atg7 to Atg3: A distinctive E1-E2 architecture and mechanism in the autophagy pathway. Mol. Cell 2011, 44, 451–461. https://doi.org/10.1016/j.molcel.2011.08.034.
- 9.
Kauffman, K.J.; Yu, S.; Jin, J.; et al. Delipidation of mammalian Atg8-family proteins by each of the four ATG4 proteases. Autophagy 2018, 14, 992–1010. https://doi.org/10.1080/15548627.2018.1437341.
- 10.
Agrotis, A.; von Chamier, L.; Oliver, H.; et al. Human ATG4 autophagy proteases counteract attachment of ubiquitin-like LC3/GABARAP proteins to other cellular proteins. J. Biol. Chem. 2019, 294, 12610–12621. https://doi.org/10.1074/jbc.AC119.009977.
- 11.
Nguyen, T.N.; Padman, B.S.; Zellner, S.; et al. ATG4 family proteins drive phagophore growth independently of the LC3/GABARAP lipidation system. Mol. Cell 2021, 81, 2013–2030.e9. https://doi.org/10.1016/j.molcel.2021.03.001.
- 12.
Jia, J.; Wang, F.; Bhujabal, Z.; et al. Stress granules and mTOR are regulated by membrane Atg8ylation during lysosomal damage. J. Cell Biol. 2022, 221, e202207091. https://doi.org/10.1083/jcb.202207091.
- 13.
Carosi, J.M.; Nguyen, T.N.; Lazarou, M.; et al. ATG8ylation of proteins: A way to cope with cell stress? J. Cell Biol. 2021, 220, e202108120. https://doi.org/10.1083/jcb.202108120.
- 14.
Ketteler, R.; Kiso, K.; von Chamier, L.; Agrotis, A. ATG5 is dispensable for ATG8ylation of cellular proteins. Autophagy Rep. 2024, 3, 2392450. https://doi.org/10.1080/27694127.2024.2392450.
- 15.
Xie, H.; Long, J.; Huang, Y.; et al. Identification of ATG7 as a protein ATG8ylation substrate. Cell Rep. 2025, 44, 116485. https://doi.org/10.1016/j.celrep.2025.116485.
- 16.
Torres, P.; Rico-Rios, S.; Ceron-Codorniu, M.; et al. TDP-43 regulates LC3ylation in neural tissue through ATG4B cryptic splicing inhibition. Acta Neuropathol. 2024, 148, 45. https://doi.org/10.1007/s00401-024-02780-4.

This work is licensed under a Creative Commons Attribution 4.0 International License.