Using network pharmacology, we identified the active ingredients and potential targets of the “Danshen & Hawthorn” herb pair for treating NAFLD. Disease targets for NAFLD were retrieved from relevant databases, and a “herb-ingredient-target” network was constructed. KEGG pathway analysis revealed that the therapeutic mechanism was associated with the IL-17, TNF, and PPAR signaling pathways, among others. Experimental validation in both in vitro (cellular model of steatosis) and in vivo (zebrafish model of NAFLD) models demonstrated that the herbal extract alleviated high-fat-induced damage. Virtual molecular docking of the top-ranked compound-target pairs identified potential key binding residues: PHE 207, TYR 134, HIS 25, GLY 143, and SER 53 for Pparg-Quercetin; ILE 317, MET 355, and PHE 273 for Ppara-Eburicoic acid; and ILE 354, PHE 351, TYR 314, SER 280, and HIS 440 for Hmox1-Dehydroeburicoic acid. In conclusion, the “Danshen & Hawthorn” herb pair ameliorates NAFLD by modulating the expression of PPARα, TNF-α, and IL-17, thereby reducing inflammation and lipid accumulation. This study provides molecular-level insights by identifying critical amino acid residues for target engagement.
- Open Access
- Article
Mechanism of Action and Target Detection of Danshen & Hawthorn in the Treatment of Non-Alcoholic Fatty Liver Disease Based on Network Pharmacology and Virtual Docking Simulation
- Zirong Zhou 1,†,
- Wenxia Tang 1,†,
- Qinghao Jiao 1,
- Yaxing Li 2,
- Ying Yang 3,
- Hui Li 3,
- Jie Yu 1,*,
- Songsong Wang 3,*
Author Information
Received: 17 Oct 2025 | Revised: 06 Dec 2025 | Accepted: 14 Dec 2025 | Published: 15 Apr 2026
Abstract
Keywords
Danshen Hawthorn | NAFLD | molecular docking | molecular dynamics simulation | inflammation
1. Introduction
Non-alcoholic fatty liver disease (NAFLD) is a common liver condition with a global prevalence of 25%, and it is a leading cause of cirrhosis and hepatocellular carcinoma. Regarding mortality risk, cardiovascular disease stands as the primary cause of death, posing twice the fatal risk of liver disease itself, which ranks only third. Moreover, the disease’s epidemiological trajectory is generating a significant and escalating economic burden. The core pathogenesis of NAFLD involves the liver producing lipotoxic substances when overloaded with metabolic substrates—particularly fatty acids and fructose—triggering a complex cascade of hepatocyte stress, inflammation, and fibrosis [1]. NAFLD encompasses a disease spectrum ranging from simple steatosis (with or without mild inflammation) to non-alcoholic steatohepatitis (NASH). NASH is characterized by necroinflammation and a more rapid progression to fibrosis compared to simple steatosis [2]. In China, the prevalence of NAFLD is between 20% and 30% and continues to increase annually [3]. The pathogenesis of NAFLD is complex, making it difficult to achieve satisfactory therapeutic outcomes with conventional single-drug regimens. Owing to the complex pathogenesis of NAFLD, the efficacy of monotherapy targeting a single pathway is limited. Combination therapy (such as the combination of GLP-1 agonists, FXR agonists and ACC inhibitors) has been demonstrated to be significantly more effective than any single therapy. Concurrently, research within the field of traditional Chinese medicine has demonstrated that compound formulations comprising Chinese herbal medicines exhibit certain therapeutic advantages in treating non-alcoholic fatty liver disease (NAFLD), including but not limited to the synergistic effects of multiple components and multi-targeted actions inherent to these formulations [4]. The combination of Salvia miltiorrhiza and Hawthorn berries originates from Shi Jinmo’s Drug Pairs. Salvia miltiorrhiza, acting to “promote blood circulation and resolve stasis”, is paired with Hawthorn berries, which “aid digestion, eliminate food stagnation, regulate qi, and disperse stasis”. According to traditional Chinese medical theory, the primary pathological sites of NAFLD lie in the liver and spleen, with long-term progression affecting the kidneys. The fundamental pathology involves damage to the liver, spleen, and kidneys, while phlegm-dampness and stasis represent secondary manifestations. This leads to impaired distribution of qi, blood, and body fluids, metabolic dysfunction, and the accumulation of endogenous phlegm-dampness and stasis obstructing the liver and spleen, ultimately causing organ dysfunction. Modern pharmacological research has demonstrated the therapeutic efficacy of Salvia miltiorrhiza and hawthorn in treating NAFLD.
Salvia miltiorrhiza Bge. is a classic herb in traditional Chinese medicine for promoting blood circulation and resolving blood stasis [5]. Modern research has confirmed its significant ameliorative effect on non-alcoholic fatty liver disease [6]. Tanshinone-type compounds in Salvia miltiorrhiza (such as Tanshinone IIA) have been demonstrated to be potent activators of the transcription factor TFEB. Activation of TFEB significantly promotes lysosomal biogenesis and lipophagocytosis in hepatocytes, thereby accelerating lipid degradation. Furthermore, this pathway is closely associated with regulating gut microbiota homeostasis and improving systemic insulin resistance [7]. Salvianolic acid components (such as salvianolic acid A) mainly regulate liver lipid metabolism, improve mitochondrial function, and inhibit ferroptosis by activating the AMPK-IGFBP1 signaling axis, thereby alleviating liver injury and steatosis [8]. In addition to in-depth research on specific components, the overall efficacy evaluation indicates that the water extract of Salvia miltiorrhiza can comprehensively improve liver inflammation, steatosis and fibrosis in NAFLD models, and its effect is related to reducing oxidative stress levels and regulating multiple targets such as PPARα and MMP2 [9]. Hawthorn (Crataegus oxyacantha) is a traditional plant that is both edible and medicinal. Its extracts have been widely used in the prevention and treatment of cardiovascular diseases and metabolic syndrome [10]. In modern pharmacological research, it has been found that hawthorn can intervene in NAFLD in multiple ways. The active component hawthorn acid in hawthorn can activate the AMPK/mTOR/ATG1 signaling pathway, enhance Lipophagy function, and effectively reduce lipid accumulation in the liver of mice induced by high-fat diet. Meanwhile, the study also found that hawthorn acid could significantly reduce the levels of pro-inflammatory factors (TNF-α, IL-1β) and cell apoptosis, demonstrating direct anti-inflammatory and anti-apoptotic effects [11]. In addition, the ethanol extract of hawthorn can optimize the intestinal flora by regulating the mechanism of the “gut-liver axis”, increase the abundance of beneficial bacteria such as Lactobacillus and Lachnospiraceae, promote the production of short-chain fatty acids (SCFAs), and improve intestinal barrier function [12]. Some studies have also shown that hawthorn has a protective effect on liver fibrosis that forms in the advanced stage of NAFLD. Hawthorn herbal preparations can significantly reduce collagen deposition (Collegen I/III) and hepatic stellate cell activation markers (α-SMA) in rat models of liver fibrosis induced by CCl4. Its mechanism of action is closely related to the inhibition of oxidative stress (reducing MDA and P. carbonyl, and increasing SOD activity) and the down-regulation of the expression of key pro-fibrotic and inflammatory factors (TGF-β, NF-κB, TNF-α, IL-1β) [13]. To sum up, Salvia miltiorrhiza may focus on lipid metabolism and liver damage, while hawthorn may focus on inflammation and fibrosis. However, most current studies have focused on Danshen or hawthorn, and limited attention has been paid to the synergistic use of these two drugs.
This study aims to systematically investigate the therapeutic mechanism of the “Danshen & Hawthorn” herb pair against NAFLD. Based on network pharmacology, we identified its active ingredients, core targets, and potential signaling pathways. Molecular docking was further utilized to simulate the interactions between key compounds and targets. Additionally, the anti-NAFLD efficacy of the “Danshen & Hawthorn” extract was experimentally validated in vitro. Our integrated strategy provides a scientific basis for understanding its molecular mechanism and offers experimental evidence to support its future development and application.
2. Materials and Methods
2.1. Experimental Drugs
Danshen and Hawthorn were purchased from Soyu Civil Pharmacy, China. Danshen Batch No.: 240406, Hawthorn Batch No.: 240501.
2.2. Animals and Cell Lines
The mouse normal hepatocyte cell line AML-12 was purchased from Procell Life Science & Technology Co., Ltd. (Wuhan, China). AB strain zebrafish were obtained from the China Zebrafish Resource Center (CZRC, Wuhan, China) and maintained in the aquatic facility of the School of Pharmacy at Shandong First Medical University (Shandong, China). All experimental procedures were conducted in accordance with the NIH Guide for the Care and Use of Laboratory Animals and were approved by the Animal Ethics Committee of the university (Approval No. S6033).
2.3. Main Experimental Reagents
DMEM/F12 basal medium (Wuhan Pricella Biotechnology Co., Ltd. (Wuhan, China), Lot No. PM150312); 0.5% insulin-transferrin-selenium additive (Wuhan Pricella Biotechnology Co., Ltd., Lot No.: PB180429); dexamethasone (AbMole (Shanghai, China), Lot No. M2176); penicillin-streptomycin mixed solution (Wuhan Pricella Biotechnology Co., Ltd., Batch No.: PB180120); fetal bovine serum (Scitecher, Batch No.: C-FBS-500); oleic acid (OA, Shanghai Yuanye Biotechnology Co., Ltd. (Shanghai, China), Batch No.: S25244); palmitic acid (PA, Shanghai Yuanye Biotechnology Co., Ltd., Batch No.: B21705); bovine serum albumin (BSA, Shanghai Yuanye Biotechnology Co., Ltd., Batch No.: V34376); SPARKeasy Cellular RNA Rapid Extraction Kit (Shandong Sikejie Biotechnology Co., Ltd. (Rizhao, China), Batch No.: AA1001-B); Triglyceride (TG) Kit (NanJing JianCheng Bioengineering Institute (Nanjing, China), Batch No.: 20250610); Total Cholesterol (TC) Kit (NanJing JianCheng Bioengineering Institute, Batch No.: 20250612); BCA Protein Assay Kit (Beyotime Biotechnology, Ltd., Lot: P0012); PBS Buffer (Shandong Sikejie Biotechnology Co., Ltd., Lot: CR0014-500ML); Egg Yolk Powder (Shanghai Yuanye Biotechnology Co., Ltd., Lot: M30HS183343); Cholesterol (Shanghai Yuanye Biotechnology Co, Ltd., Lot:L26N11F132572); SPARKscript II RT Plus Kit (Shandong Sikejie Biotechnology Co., Ltd., Lot: AG0304); 2 × SYBR Green Qpcr Mix (Shandong Sikejie Biotechnology Co., Ltd., Lot: AH0104) SPARKeasy Cell RNA Rapid Extraction Kit (Shandong Sikejie Biotechnology Co., Ltd., Lot: FFEKZ).
2.4. Main Instruments
Multi-function enzyme labeller (Molecular Devices (San Jose, CA, USA), model: SpectraMaxABS); Fluorescence quantitative PCR instrument (Bio-Rade (Hercules, CA, USA), model: Roche LightCycler 480); Inverted fluorescence microscope (Sunny Optical Technology Co., Ltd. (Hong Kong, China), model: ICX41RFL series); Sample freezing and grinding instrument (Shandong, China) Ltd., model: B0-IR).
2.5. Network Pharmacological Analyses
The active ingredients of “Danshen & Hawthorn” were first retrieved from the TCM Systems Pharmacology Database and Analysis Platform (TCMSP). Screening was performed using oral bioavailability (OB ≥ 30%) and drug-likeness (DL ≥ 0.18) as criteria. This initial dataset was further supplemented with information from the Integrative Pharmacology-based Research Platform of TCM (TCMIP) and a review of relevant literature. The finalized list of active ingredients was submitted to the Swiss Target Prediction database to forecast compound targets. Finally, the predicted targets were cross-referenced and integrated with those directly obtained from the TCMSP and TCMIP databases to construct a comprehensive target list.
2.6. Establishment of NAFLD Disease Targets
Potential therapeutic targets associated with nonalcoholic fatty liver disease (NAFLD) were identified by searching the OMIM and GeneCards databases using “nonalcoholic fatty liver disease” as the keyword. The resulting target lists from both databases were then merged, and duplicate entries were removed to generate a consolidated set of NAFLD-related targets.
2.7. Constructing Protein-Protein (PPI) Interaction Network Diagrams
The drug-disease common targets were imported into the STRING database to construct a protein-protein interaction (PPI) network. The analysis was configured with the biological species set to “Homo sapiens”, a minimum required interaction score of 0.4, and all other parameters kept at their default settings.
2.8. Gene Ontology (GO) Annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Analysis
The overlapping gene targets of “Danshen & Hawthorn” associated with NAFLD were submitted to the Metascape database for Gene Ontology (GO) enrichment analysis to identify significantly enriched biological processes. Significantly enriched entries are defined as those with an FDR (Benjamini-Hochberg correction) < 0.05. The background gene set utilizes Metascape’s default Homo sapiens genome-wide background.
2.9. Virtual Docking Screening of Detection Sites
The three-dimensional structures of the core targets were obtained from the Protein Data Bank (PDB), while the structures of the small molecule compounds were retrieved from the PubChem database. The molecular docking program was carried out using Discovery Studio 2019. The Prepare Protein in the Macromolecules module was utilized to pretreat the proteins (including dewatering, hydrogenation, energy minimization, etc.). The Small Molecules were pretreated using Prepare Ligands in the Small Molecules section. Then use the receptor-ligand Interactions plate to select the Define Receptor to define the protein, and the spatial coordinates utilize the From Receptor Cavities in the Define Site. Select the default first spatial coordinate based on the protein’s own spatial structure, and modify the coordinate range Radius to 10. Select Dock Ligands (Libdock) in Docking mode, select User Specified in Docking Preferences, set Max Hits to Save to 5, all default Settings unless otherwise specified. Finally, for each protein and ligand, the docking result ranked first according to the Libdock Score was taken for integration. Molecular dynamics simulations were performed using the Schrodinger Maestro software. Protein Preparation was used to enter the protein preparation, and Fill in Missing Loops was checked in the Preprocess to ensure the integrity and reliability of the 3D protein model. Subsequently, the System was established through System Build, the water environment was selected by Predefined to select TIP3P, and the Minimize Volume energy minimization simulation box. Select Neutralize by Adding in the ions interface and confirm Recalculate. At the same time, set Add Salt, Cl−, Na+, 0.15 M and other parameters by default. After the simulation system was established, the related Settings of molecular dynamics simulation were carried out. The simulation time was 100 ns, the parameter Recording interval related Trajectory was set to 100, the Energy was set to 1.2, and the Ensemble class was set to NPT. The Temperature (K) is 300 K, and the Pressure (bar) is 1.01325 bar. Alanine scanning was performed using Discovery Studio 2019. After molecular docking, all amino acids within a radius of 3 Å centered on the ligand were selected as mutation points. The identification of key alanine residues was similar to that of molecular docking. The final key residues were determined by the combination of molecular docking, molecular dynamics simulation and alanine scanning.
2.10. Preparation of Danshen Hawthorn Extract
The “Danshen & Hawthorn” extract was prepared from a mixture of 15 g of Danshen and 15 g of hawthorn. The herbs were subjected to ultrasonic-assisted extraction with 600 mL of 70% ethanol at 80% power and 60 °C for 1 h. The resulting extract was concentrated to a dense residue using a rotary evaporator, dried at 37 °C, and stored at 4 °C. For bioassays, 30 mg and 60 mg aliquots of the dried extract were dissolved in 1 mL of purified water, respectively. The solutions were sterilized by filtration through a 0.22 μm membrane and stored at −20 °C until use.
2.11. Cellular Modelling Drug and Zebrafish Modelling Drug Configuration
40% BSA: 0.6 g of BSA was dissolved in 1.5 mL of PBS pre-warmed to 55 °C with gentle mixing. 20% BSA: 0.3 g of BSA was dissolved in 1.5 mL of PBS pre-warmed to 55 °C with gentle mixing. 0.1 M NaOH: 0.04 g of NaOH solid was dissolved in 10 mL of purified water. 40 mM Oleic Acid (OA): 19.04 μL of OA was mixed with 1.5 mL of 0.1 M NaOH and saponified in a 75 °C water bath. The solution was stored at −20 °C. 20 mM Palmitic Acid (PA): 0.007675 g of PA was mixed with 1.5 mL of 0.1 M NaOH, saponified in a 75 °C water bath for 30 min, and stored at −20 °C. DMEM/F12 Complete Medium: 500 mL of DMEM/F12 basal medium was supplemented with 10% FBS, 0.5% ITS-G, 40 ng/mL dexamethasone, and 1% penicillin-streptomycin. The medium was sterile-filtered, thoroughly mixed, and stored at 4 °C. Preparation of high-fat diet for zebrafish: 600 mg of cholesterol was dissolved in 40 mL of absolute ethanol using ultrasonication. This solution was then gradually added to 11.4 g of egg yolk powder in a mortar while grinding continuously. The mixture was evaporated in a 60 °C water bath to remove the absolute ethanol. The high-fat diet was fully prepared once the ethanol had completely evaporated.
2.12. Cellular Drug Delivery and Related Indicator Assays
The culture media for each group were prepared in 10% complete medium. The blank control group was treated with 3 mL of medium supplemented with 3 μL of 20% BSA and 3 μL of 40% BSA. The model group was treated with 3 mL of medium supplemented with 3 μL of oleic acid (OA) and 3 μL of palmitic acid (PA). The two drug treatment groups were treated with 3 mL of medium containing the same concentrations of OA and PA, further supplemented with 3 μL of Salvia miltiorrhiza and hawthorn extract at either 30 mg/mL or 60 mg/mL, respectively. Twenty-four hours after treatment, intracellular levels of total cholesterol (TC) and triglycerides (TG) were measured. Concurrently, the mRNA expression levels of PPARα, TNF-α, and IL-17 were determined using quantitative real-time PCR. All assays were strictly performed following the manufacturers’ protocols.
2.13. Zebrafish Drug Administration and Related Indicator Tests
Juvenile AB strain zebrafish at 5 days post-fertilization (dpf) were distributed into six-well plates, with 30 fish per well, for modeling and drug treatment. The fish were randomly divided into the following groups: a blank group (fed with egg yolk powder), a model group (fed with a 5%cholesterol high-fat diet), and treatment groups (fed with the same 5% cholesterol high-fat diet supplemented with Danshen & Hawthorn extract at 30 mg/mL or 60 mg/mL). The modeling and treatment continued for five days, during which the fish culture water was changed twice daily. At the end of the experiment, the zebrafish were fasted for 24 h. Subsequently, the whole-body levels of total cholesterol (TC) and triglycerides (TG) were measured. In parallel, the gene expression levels of PPARα, TNF-α, and IL-17 were analyzed. All measurements were conducted in strict accordance with the manufacturers’ instructions.
2.14. Statistical Analysis
All data are presented as mean ± SEM using unpaired one-way ANOVA or t-test, followed by tests for between-group or multiple recombination comparisons. A p value of less than 0.05 was considered significant. All data were analyzed using GraphPad Prism (10.1.0).
3. Results
3.1. Targets Related to the Treatment of NAFLD with Danshen & Hawthorn
Potential targets associated with nonalcoholic fatty liver disease (NAFLD) were retrieved from the OMIM and GeneCards databases using “nonalcoholic fatty liver disease” as the search term. The results from both databases were then consolidated, yielding a total of 1235 NAFLD-related targets. By integrating TCMSP and TCMIP database information, 379 potential therapeutic targets for components of Salvia miltiorrhiza and 305 potential therapeutic targets for components of hawthorn were obtained. Venn diagram results showed that there were 96 intersection targets between Salvia miltiorrhiza and NAFLD, indicating that the number of targets of Salvia miltiorrhiza in the treatment of NAFLD was 96. Similarly, there were 66 intersection targets of hawthorn and NAFLD, indicating that the number of targets of hawthorn in the treatment of NAFLD was 66. Among them, there were 27 intersection targets of Danshen and Haw and NAFLD, indicating that there were 27 targets of Danshen and Haw in the treatment of NAFLD. The result is presented in Figure 1.
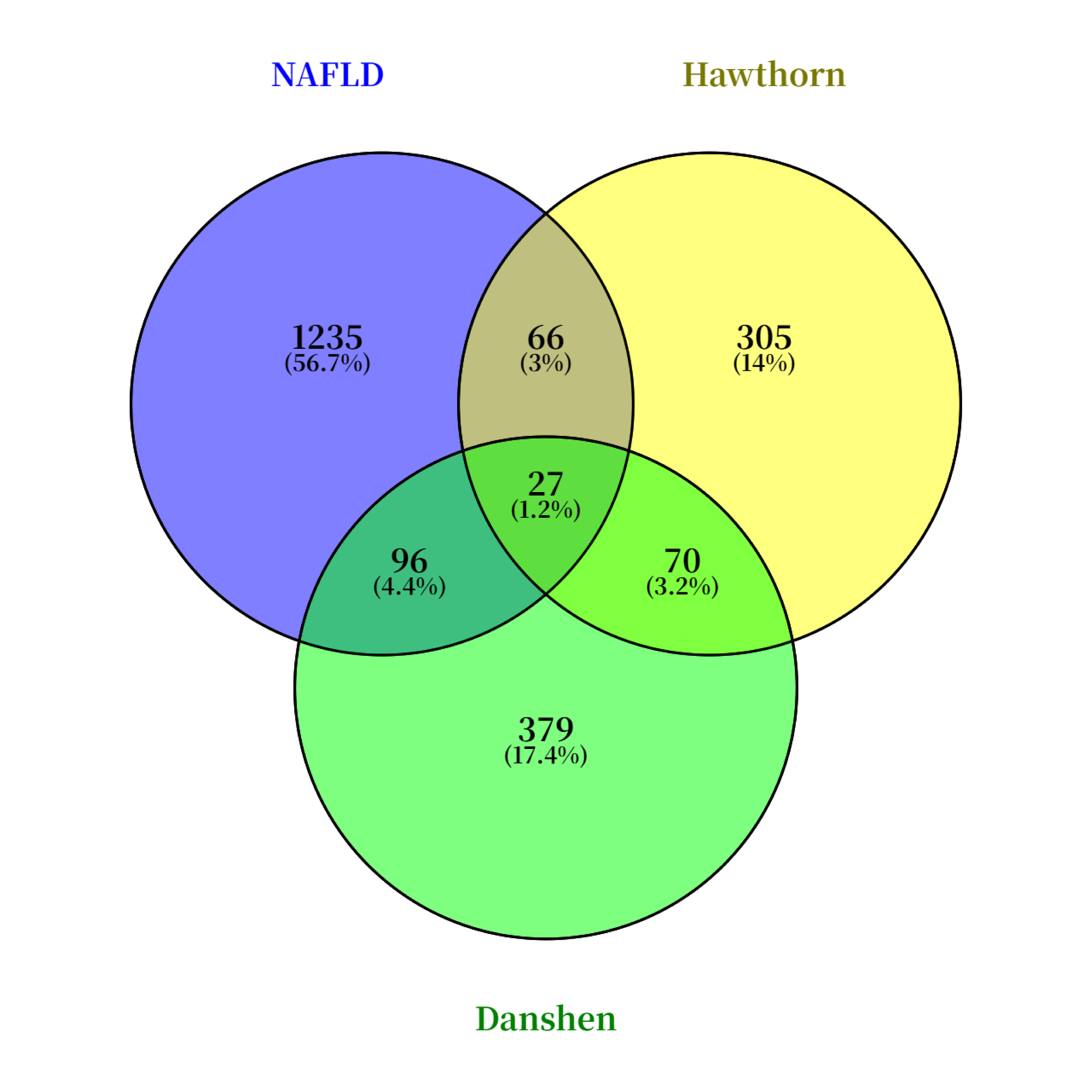
3.2. Drug-Component-Target Network Diagram Construction
Using the TCMSP and TCMIP databases, 57 active ingredients from Danshen and 47 from Hawthorn were identified, along with their corresponding targets. This information regarding the herbs, ingredients, and targets was then imported into Cytoscape software to construct the herb-ingredient-target network, as shown in Figure 2.
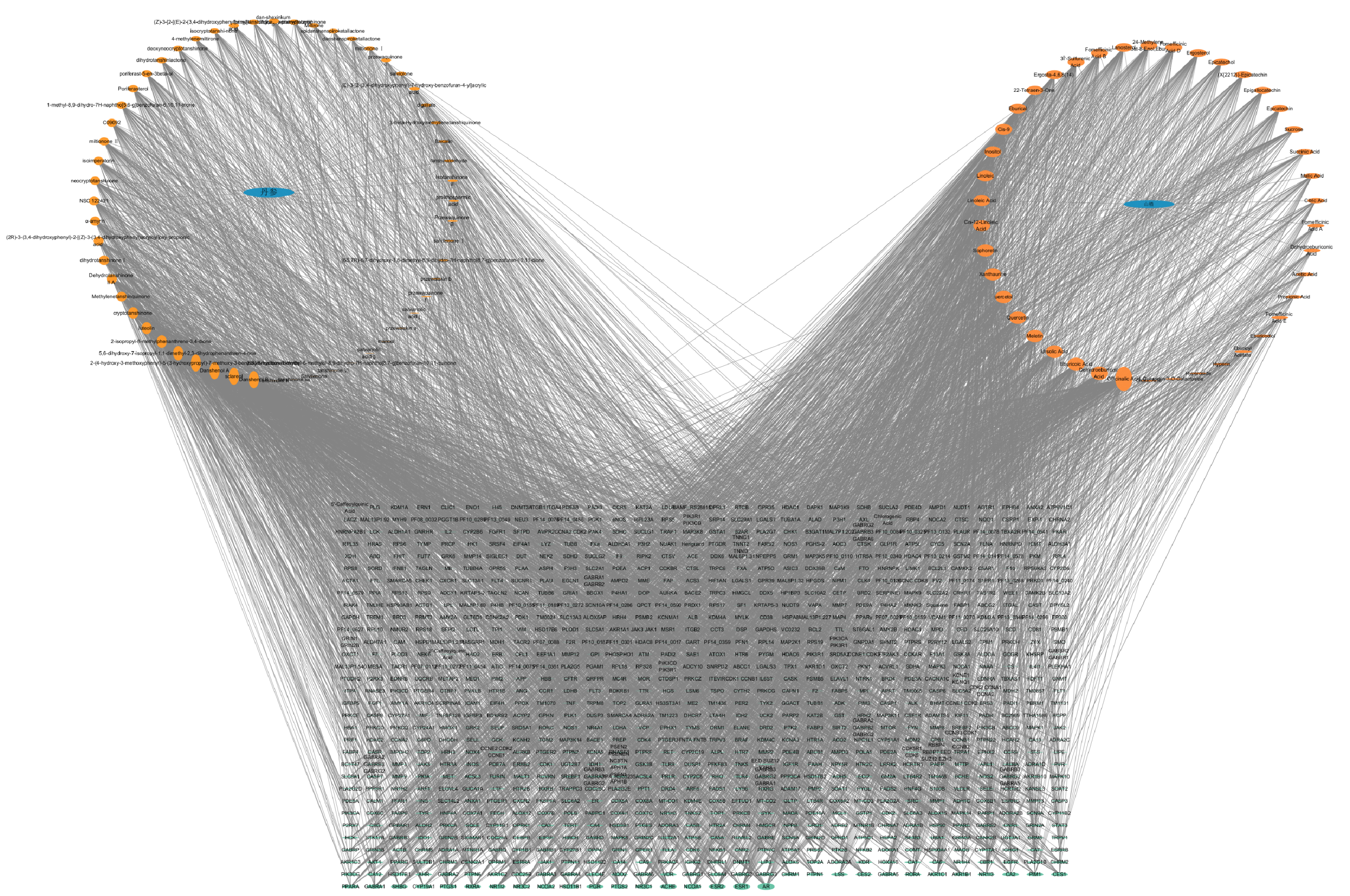
3.3. PPI Network Construction
The overlapping genes between “Danshen & Hawthorn” and NAFLD were submitted to the STRING database to generate a protein-protein interaction (PPI) network. The resulting network contained 26 nodes and 112 edges, with an average node degree of 8.62 and a local clustering coefficient of 0.62 (Figure 3). By analyzing the PPI of 26 intersection targets and then importing them into the Cytoscape network to sort them in descending order using Degree values, the top ten targets are selected as core targets for subsequent analysis, and the resulting network is displayed in Figure 4.
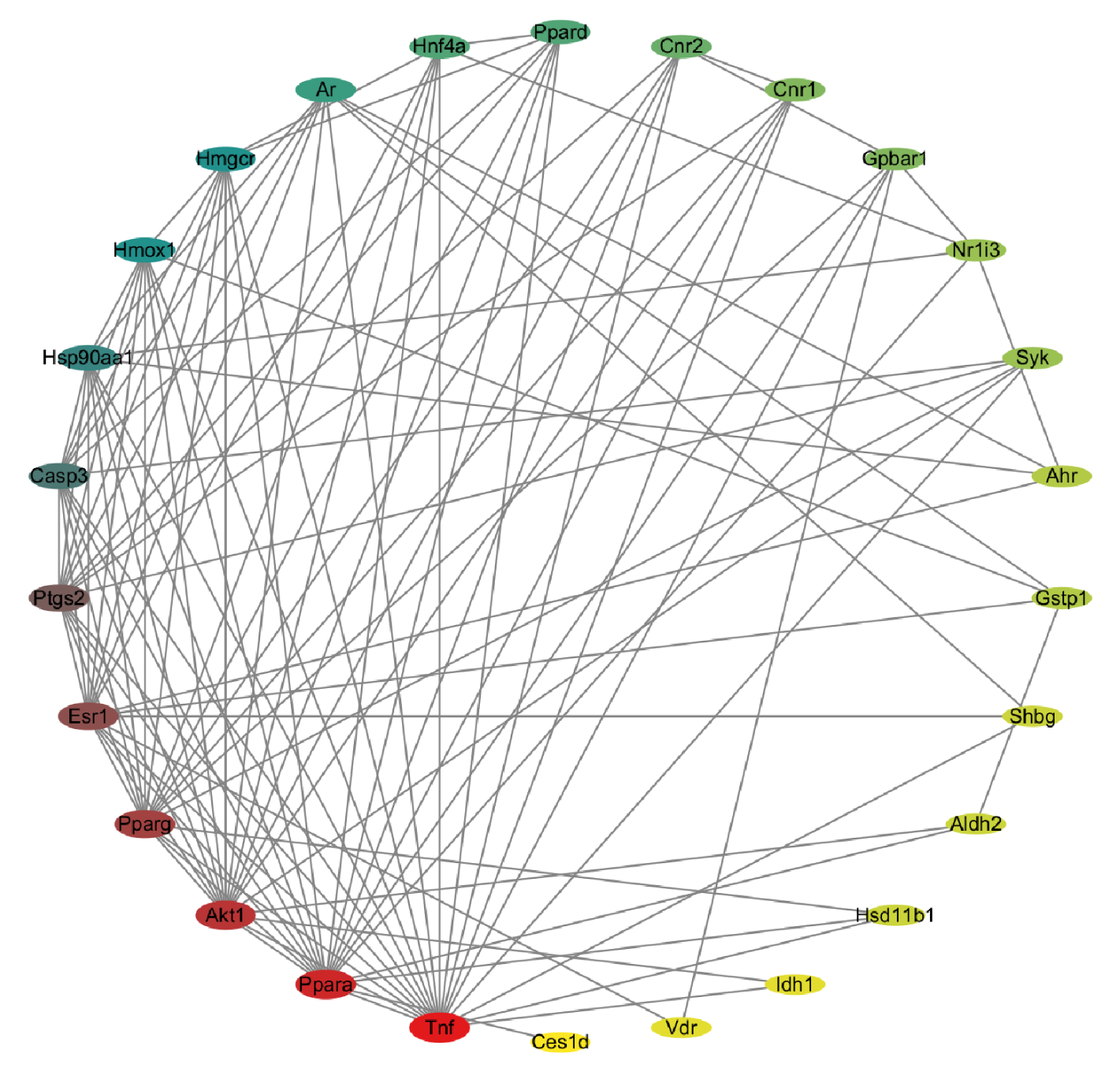
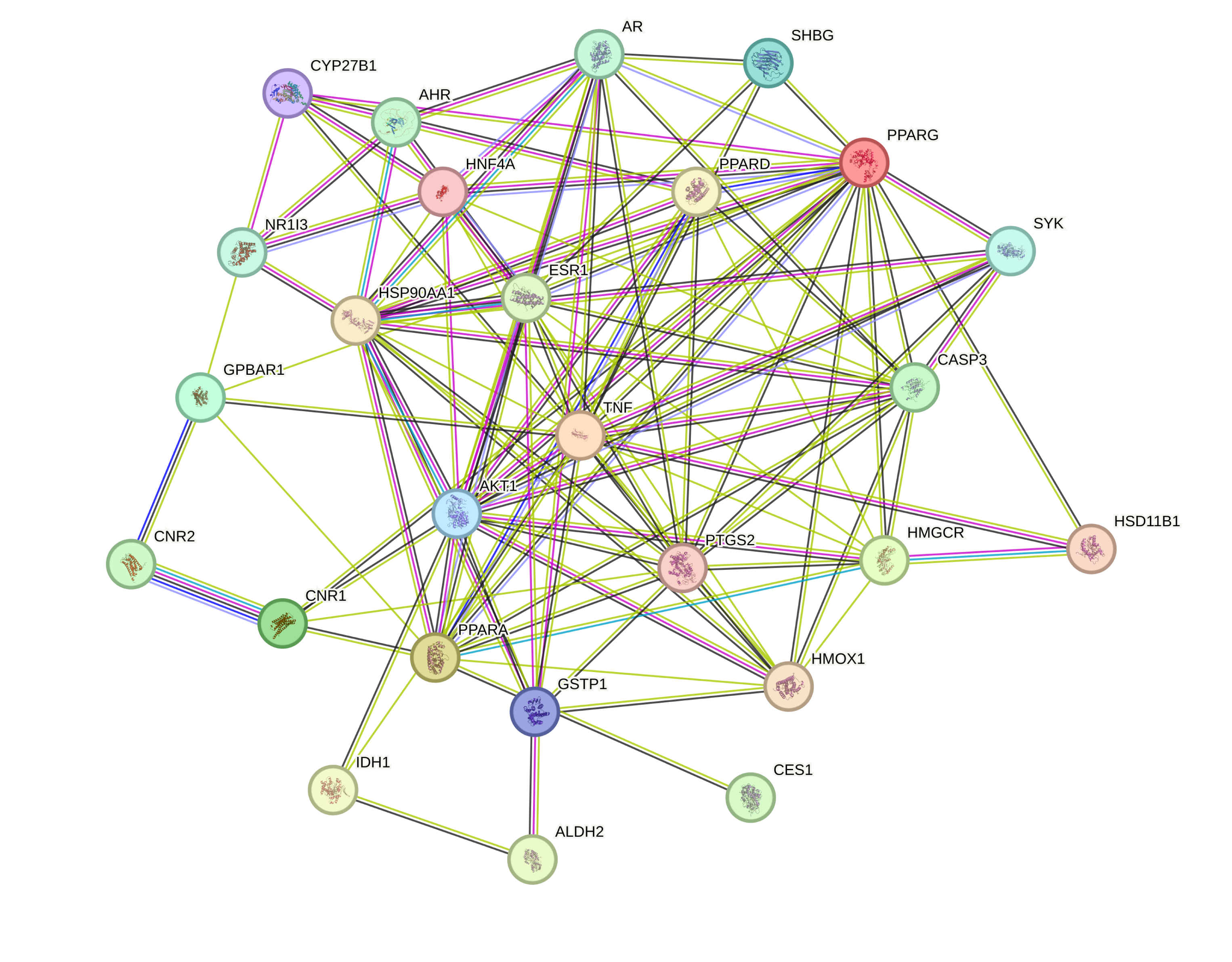

3.4. GO and KEGG Analyses
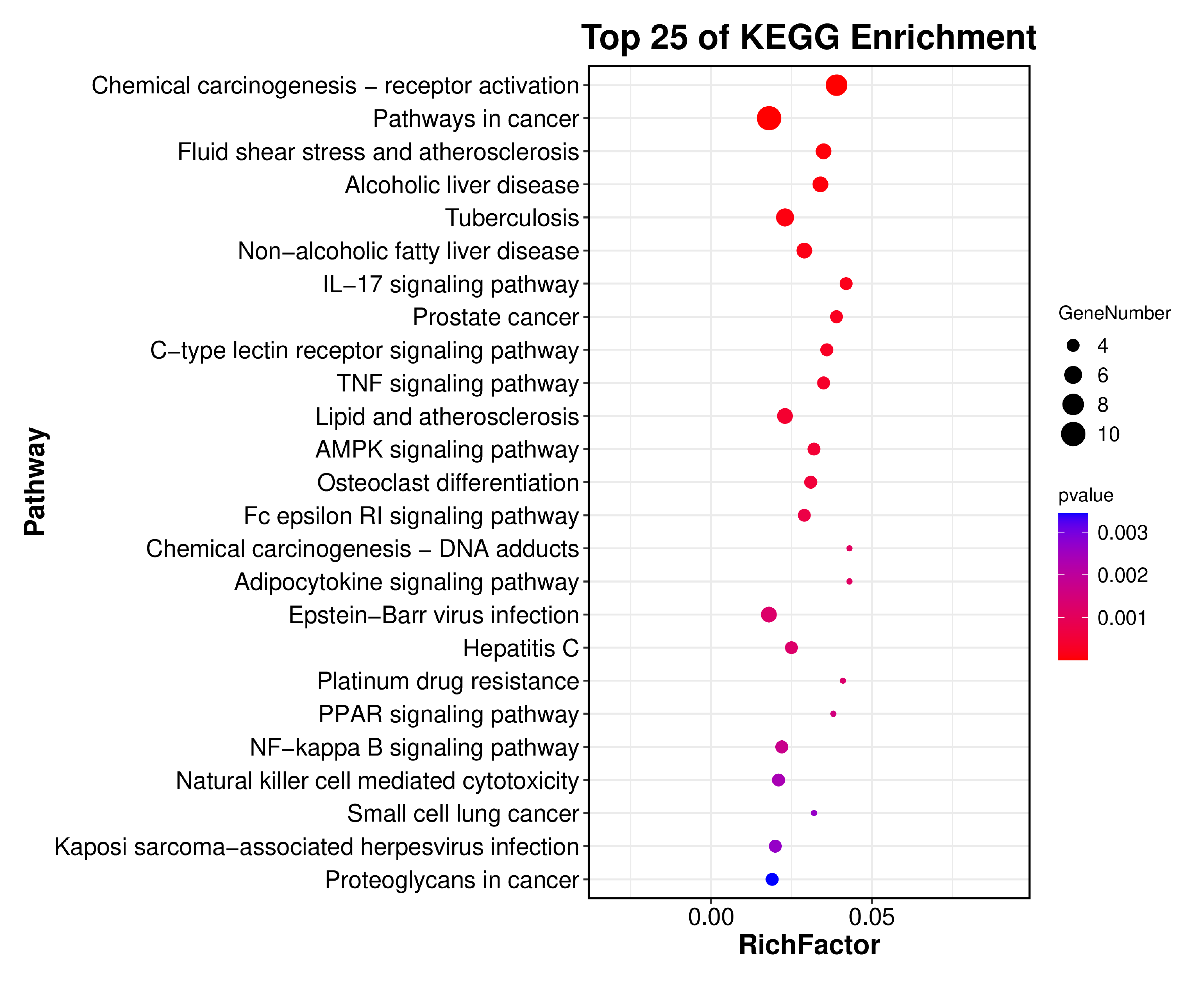
GO and KEGG pathway enrichment analyses were performed for the relevant targets using the Metascape website. The GO analysis encompassed three categories: cellular component, molecular function, and biological process. The top 25 most significant terms from the GO analysis were selected for visualization (Figure 5). Similarly, the top 25 most significant pathways from the KEGG analysis were visualized (Figure 6). GO functional enrichment analysis showed that the potential targets of Salvia miltiorrhiza and hawthorn in the treatment of NAFLD were mainly involved in biological process regulation, metabolic process, positive regulation and negative regulation of biological process, etc. The cellular component (CC) mainly involved cell anatomical entities and protein complexes. The molecular function (MF) mainly involves catalytic activity, molecular transcription activity, transcriptional regulatory activity, etc. KEGG pathway enrichment analysis showed that the potential targets were mainly involved in non-alcoholic fatty liver disease, IL-17 signaling pathway, TNF signaling pathway, lipid and atherosclerosis, PPAR signaling pathway, etc. Path selection is guiding the experiment and is not an after-the-fact rationalization.
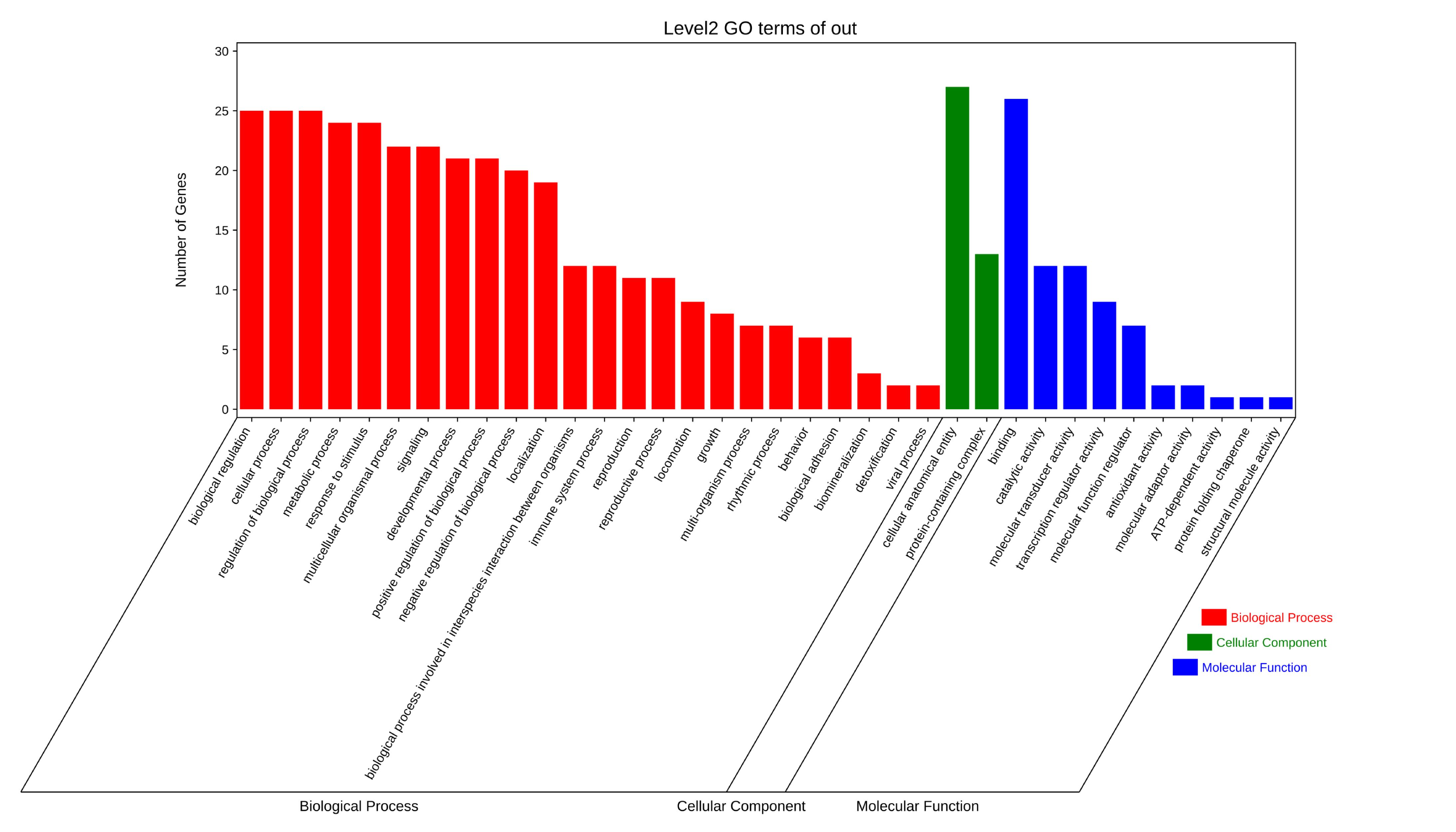
3.5. Virtual Docking Screening
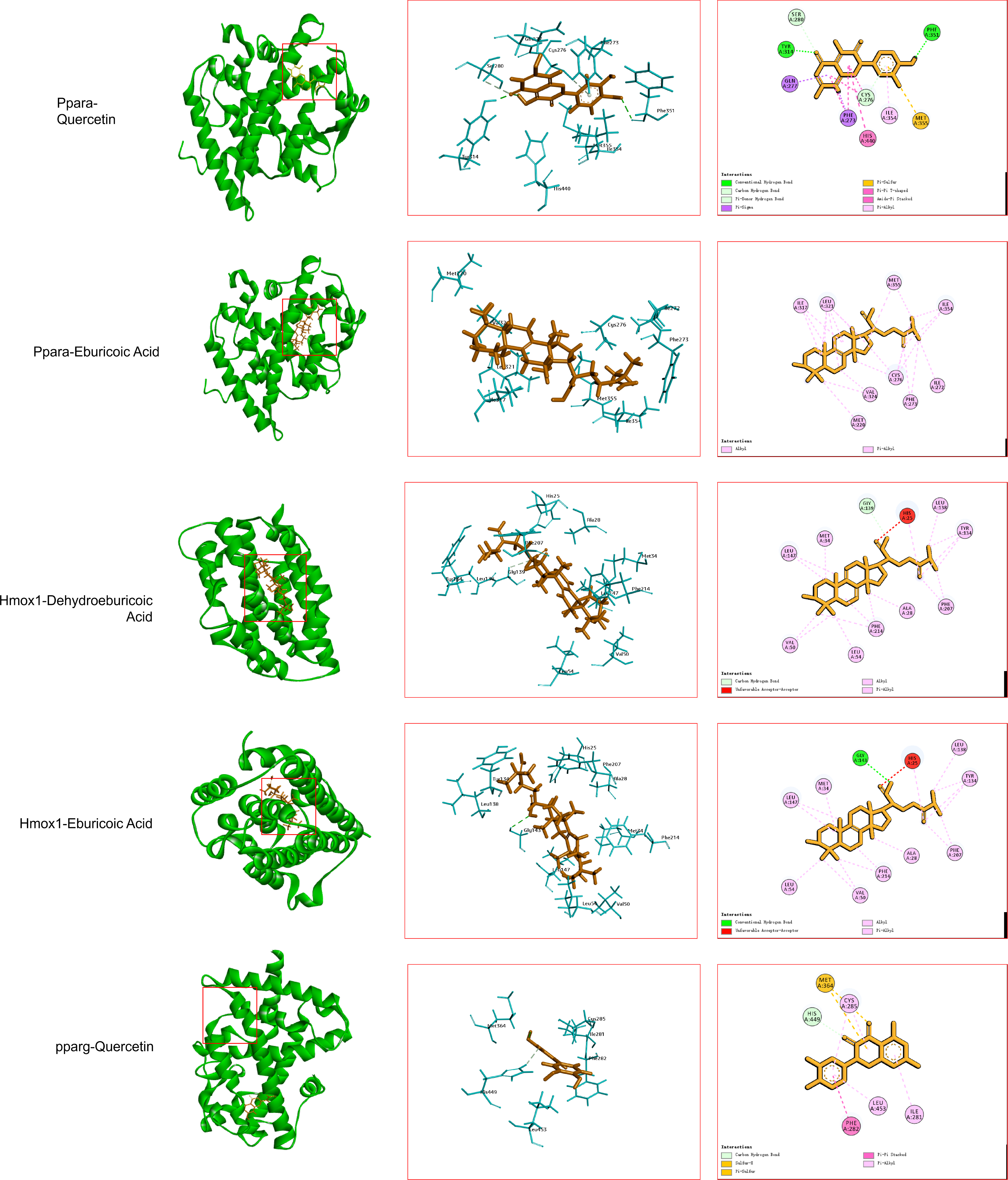
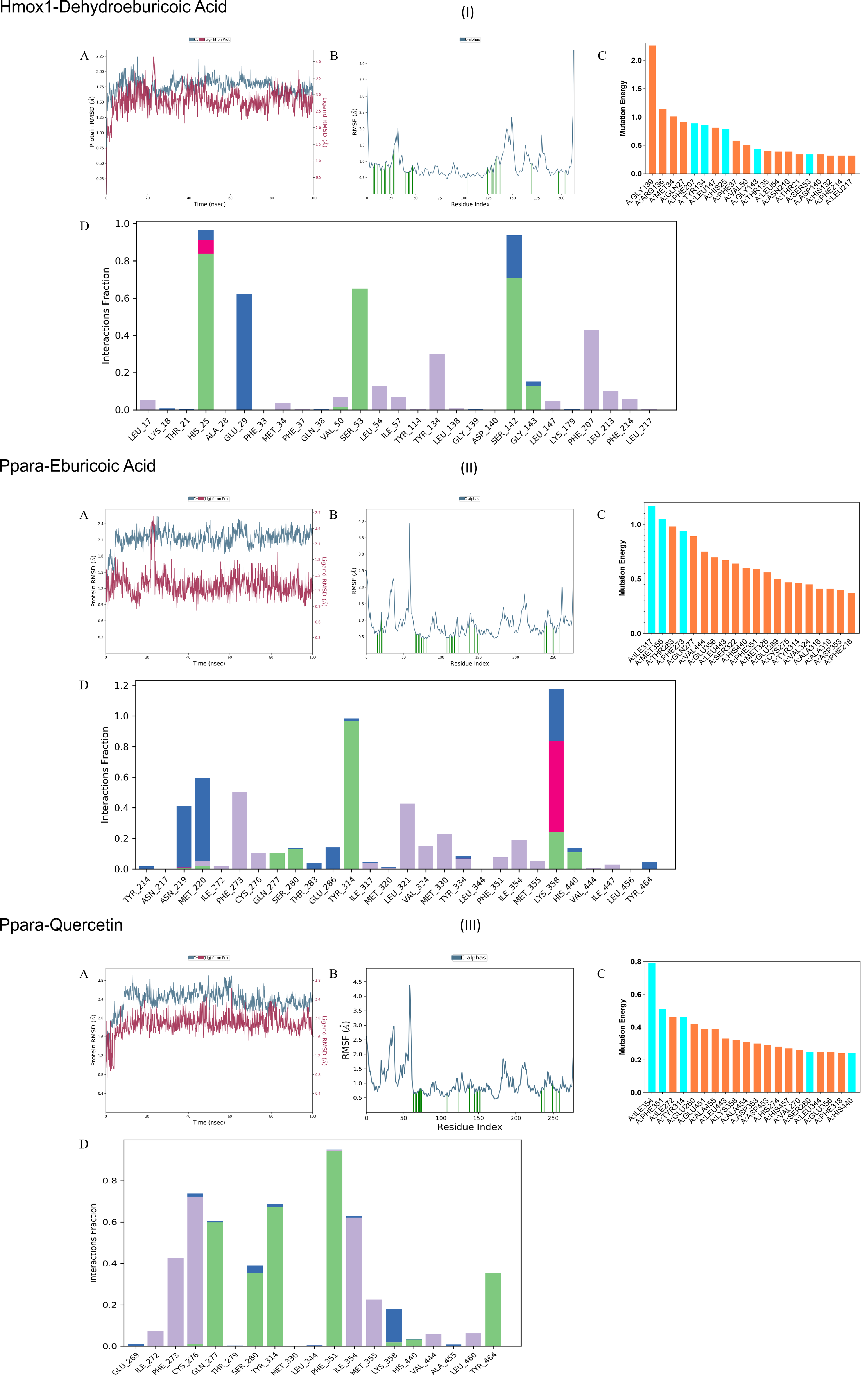
Molecular docking was performed between the screened key active components and their corresponding core targets. All the compounds were selected by the top 10 Degree value. The compounds and targets involved in the table are used to provide a basis for subsequent molecular docking (Table 1). Table 2 lists the molecular docking with the 10 targets and the molecular docking scores for the 10 compounds involved in Table 1. The top five complexes with the highest docking scores were selected for visual analysis (Figure 7). To further investigate the binding interactions, the top three ranked complexes from the docking results were subjected to molecular dynamics simulations (Figure 8). The molecular dynamic simulation mainly shows the binding degree of the ligand and the protein. RMSD shows that the three groups of protein molecules have a good binding degree. The green line in the RMSF figure contacts the RMSF, indicating the contact between the ligand and the protein at the amino acid residue. These results indicate that the ligands mainly interact in the secondary structure of the protein, and there are no flexible regions or disordered regions. Alanine scan showed that all amino acids in the range of ligand 3 Å were mutated to alanine, and the mutation energy was ranked. The alanine scan result would show “stable” or “unstable”, and “stable” indicated that the point mutation to alanine had no effect on the protein structure, and “unstable” indicated that the point mutation to alanine had an effect on the protein structure. All the results of “stable” and “unstable” were statistically analyzed, and the top 20 sites were taken as potential sites affecting the protein structure, and 20 sites were sufficient for inclusion. Meanwhile, the core residues in the ligand complex were illustrated by the interaction sites of the ligand and protein in molecular dynamic simulation and molecular docking. The integrated analysis from molecular docking, protein-ligand interaction examination, and alanine scanning revealed the following potential key residues: For PPARα-Quercetin: PHE 207, TYR 134, HIS 25, GLY 143, and SER 53. For PPARα-Eburicoic Acid: ILE 317, MET 355, and PHE 273. For HMOX1-Dehydroeburicoic Acid: ILE 354, PHE 351, TYR 314, SER 280, and HIS 440.
Top 10 key ingredients and key targets.
|
Key Ingredient |
Key Target |
|---|---|
|
Ursolic Acid |
Tnf |
|
Officinalic Acid |
Ppara |
|
Sclareol |
Akt1 |
|
Quercetin |
Pparg |
|
Meletin |
Esr1 |
|
Sophoretin |
Ptgs2 |
|
Eburicoic Acid |
Casp3 |
|
Dehydroeburicoic Acid |
Hsp90aa1 |
|
Danshenol B |
Hmox1 |
|
Danshenol A |
Hmgcr |
Molecular docking scores of the top 10 key components and key targets.
|
Protein |
Molecule |
LibDock Score |
|---|---|---|
|
AKT1 |
Ursolic Acid |
69.0701 |
|
Officinalic Acid |
75.2435 |
|
|
Sclareol |
80.394 |
|
|
Quercetin |
93.5439 |
|
|
Meletin |
93.5439 |
|
|
Sophoretin |
93.5439 |
|
|
Eburicoic Acid |
76.9732 |
|
|
Dehydroeburicoic Acid |
76.6155 |
|
|
Danshenol B |
96.8538 |
|
|
Danshenol A |
85.6697 |
|
|
Casp3 |
Sclareol |
81.5713 |
|
Quercetin |
100.065 |
|
|
Meletin |
100.065 |
|
|
Sophoretin |
100.065 |
|
|
Danshenol B |
84.2001 |
|
|
Esr1 |
Ursolic Acid |
101.046 |
|
Officinalic Acid |
100.618 |
|
|
Sclareol |
97.7842 |
|
|
Quercetin |
109.102 |
|
|
Meletin |
109.102 |
|
|
Sophoretin |
109.102 |
|
|
Eburicoic Acid |
104.524 |
|
|
Dehydroeburicoic Acid |
109.399 |
|
|
Danshenol B |
108.427 |
|
|
Danshenol A |
106.539 |
|
|
Hmgcr |
Ursolic Acid |
99.9889 |
|
Officinalic Acid |
109.309 |
|
|
Sclareol |
86.654 |
|
|
Quercetin |
84.5055 |
|
|
Meletin |
84.5055 |
|
|
Sophoretin |
84.5055 |
|
|
Eburicoic Acid |
108.816 |
|
|
Dehydroeburicoic Acid |
105.662 |
|
|
Danshenol B |
88.607 |
|
|
Danshenol A |
91.1685 |
|
|
Hmox1 |
Ursolic Acid |
101.65 |
|
Officinalic Acid |
105.266 |
|
|
Sclareol |
91.665 |
|
|
Quercetin |
95.3222 |
|
|
Meletin |
95.3222 |
|
|
Sophoretin |
95.3222 |
|
|
Eburicoic Acid |
118.024 |
|
|
Dehydroeburicoic Acid |
120.39 |
|
|
Danshenol B |
102.31 |
|
|
Danshenol A |
98.5583 |
|
|
Hsp90aa1 |
Ursolic Acid |
78.2817 |
|
Officinalic Acid |
85.5568 |
|
|
Sclareol |
78.7152 |
|
|
Quercetin |
89.1937 |
|
|
Meletin |
89.1937 |
|
|
Sophoretin |
89.1937 |
|
|
Eburicoic Acid |
89.4937 |
|
|
Dehydroeburicoic Acid |
98.0082 |
|
|
Danshenol B |
92.7388 |
|
|
Danshenol A |
91.1935 |
|
|
Ppara |
Ursolic Acid |
92.9089 |
|
Sclareol |
106.478 |
|
|
Quercetin |
126.925 |
|
|
Meletin |
126.925 |
|
|
Sophoretin |
126.925 |
|
|
Eburicoic Acid |
119.902 |
|
|
Dehydroeburicoic Acid |
117.678 |
|
|
Danshenol B |
116.612 |
|
|
Danshenol A |
115.257 |
|
|
Pparg |
Sclareol |
104.083 |
|
Quercetin |
118.802 |
|
|
Meletin |
118.802 |
|
|
Sophoretin |
118.802 |
|
|
Ptgs2 |
Sclareol |
104.019 |
|
Quercetin |
89.5135 |
|
|
Meletin |
89.5135 |
|
|
Sophoretin |
89.5135 |
|
|
Danshenol B |
96.631 |
|
|
Danshenol A |
89.653 |
|
|
Tnfa |
Sclareol |
79.7517 |
|
Quercetin |
72.7876 |
|
|
Meletin |
72.7876 |
|
|
Sophoretin |
72.7876 |
|
|
Eburicoic Acid |
77.7034 |
|
|
Dehydroeburicoic Acid |
70.2141 |
|
|
Danshenol B |
71.116 |
|
|
Danshenol A |
66.952 |
3.6. Effects of Danshen & Hawthorn Extract on OA and PA-Induced Intracellular TC and TG Content in AML-12 Cells and on Gene Expression Levels of PPARa, TNFa and IL-17
As shown in Figure 9, the intracellular levels of total cholesterol (TC) and triglycerides (TG) in AML-12 cells were significantly elevated by OA and PA induction compared to the blank group, indicating substantial lipid accumulation. Treatment with the “Danshen & Hawthorn” extract significantly reduced these elevated TC and TG levels, demonstrating its ability to ameliorate lipid accumulation induced by the high-fat model. Furthermore, the mRNA expression levels of PPARα, TNF-α, and IL-17 were measured. The administration of the Danshen-Hawthorn extract reversed the dysregulation of these genes caused by high-fat induction. The modulation of PPARα expression suggests that the extract interferes with NAFLD progression related to lipid metabolism, while the changes in TNF-α and IL-17 levels indicate an anti-inflammatory effect contributing to its therapeutic impact.
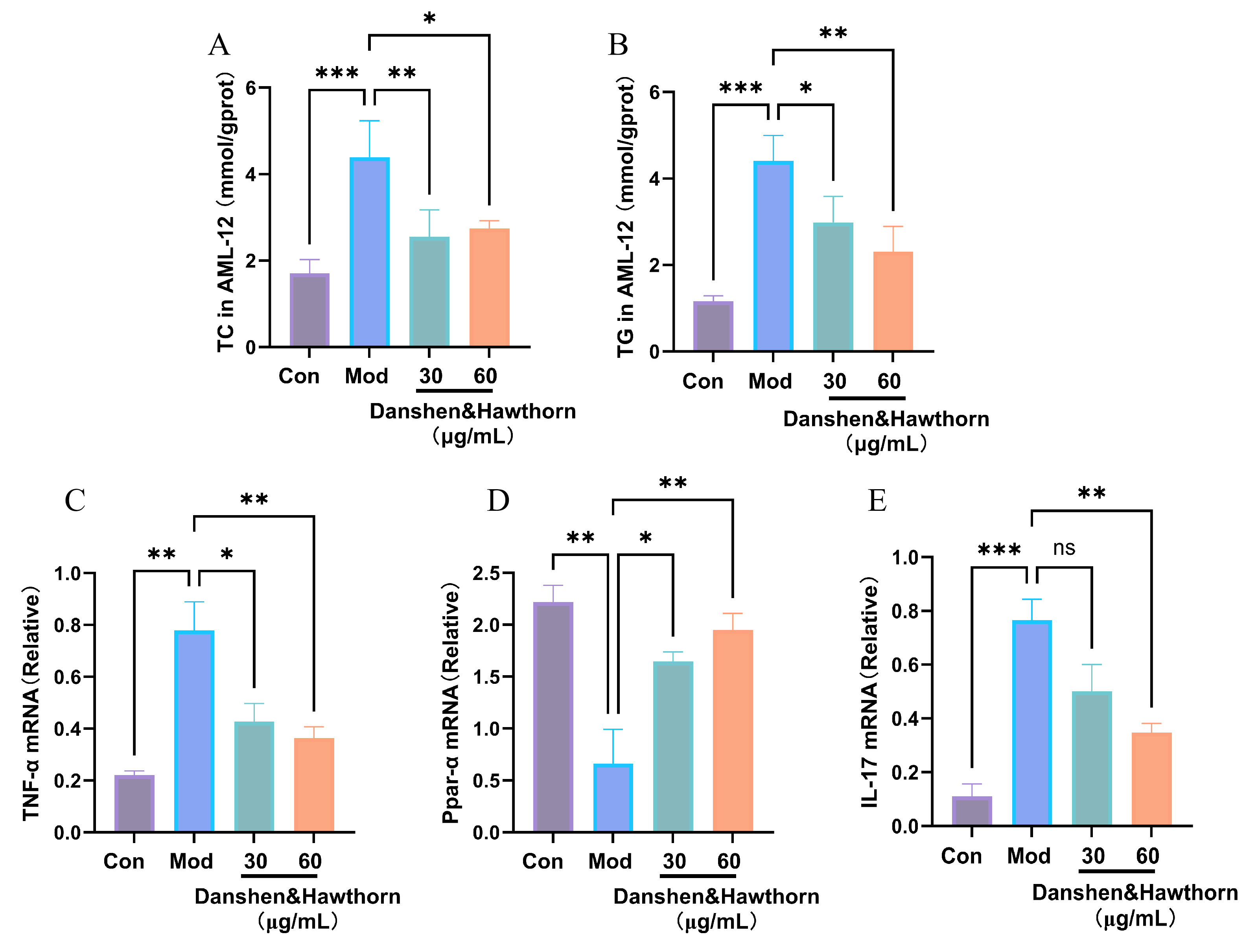
3.7. Effects of Danshen & Hawthorn Hawthorn Extract on TC and TG Content and on Gene Expression Levels of PPARa, TNFa and IL-17 in Zebrafish Induced by High-Fat Diets
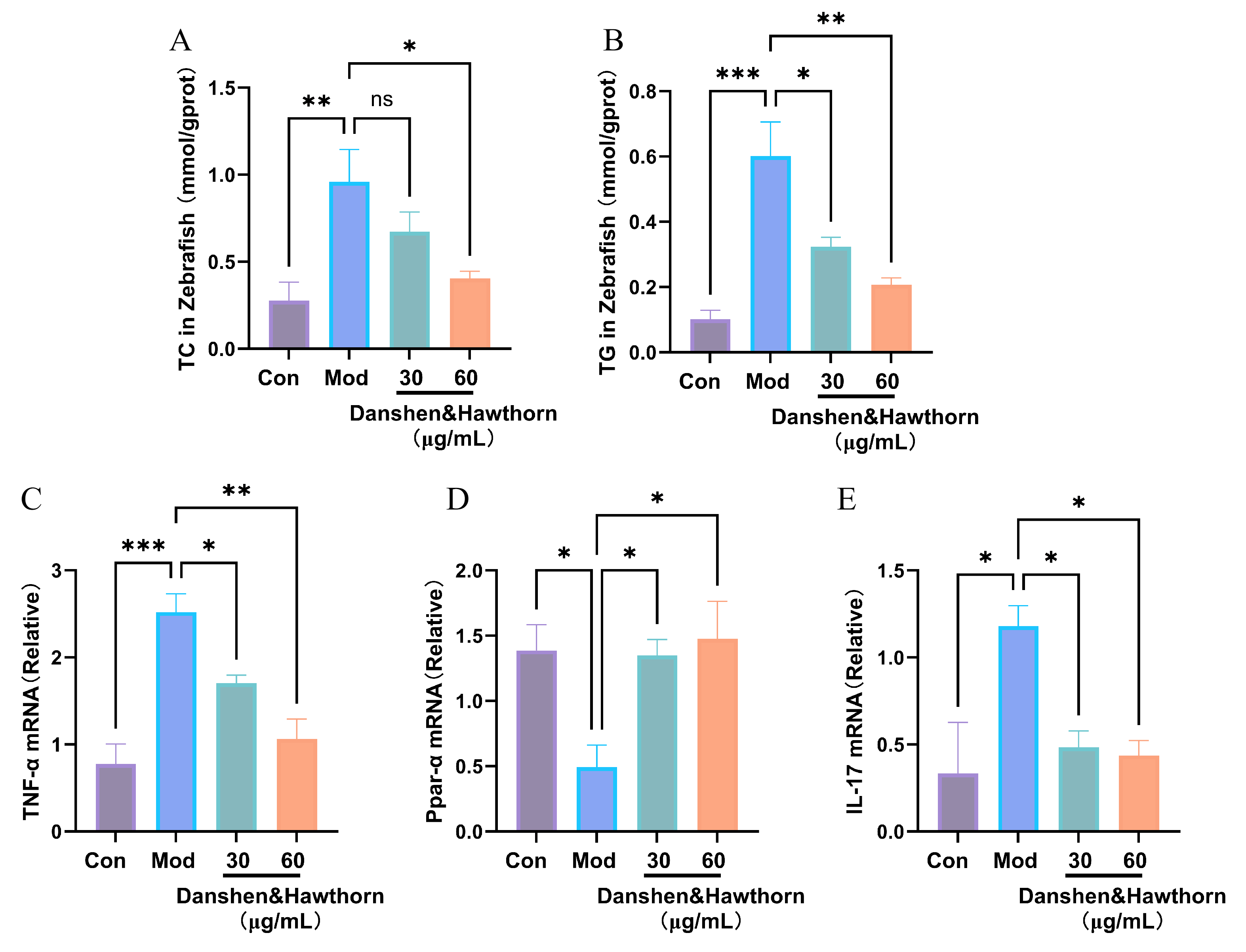
As shown in Figure 10, zebrafish exposed to a high-fat diet (HFD; 5% cholesterol) exhibited significantly elevated levels of total cholesterol (TC) and triglycerides (TG) compared to the blank group (fed with egg yolk powder), indicating successful induction of lipid accumulation. Treatment with the Danshen-Hawthorn extract significantly reduced these elevated TC and TG levels in a dose-dependent manner, demonstrating its efficacy in alleviating HFD-induced lipid accumulation in zebrafish.
Furthermore, the mRNA expression levels of PPARα, TNF-α, and IL-17 in zebrafish were analyzed. The extract administration effectively reversed the HFD-induced dysregulation of these genes. Specifically, the modulation of PPARα implies that the therapeutic effect involves lipid metabolic pathways, while the alterations in TNF-α and IL-17 levels suggest an anti-inflammatory mechanism. These findings are consistent with the observations in the cellular model.
4. Discussion
The herb pair “Danshen & Hawthorn” originates from “Shi Jinmo’s Drug Pairs”. Danshen (Salvia miltiorrhiza) is known to activate blood circulation, resolve blood stasis, regulate menstruation, and alleviate pain. When combined with Hawthorn, its power to promote blood circulation and remove stasis is significantly enhanced. This combination is effective in treating cardiovascular diseases and nodules caused by blood stasis and obstruction. It also aids in lowering blood pressure and lipids, and provides hepatoprotective effects. As a prominent traditional Chinese medicine, Danshen has a long history of use in treating blood stasis syndrome and related disorders [5]. Its active components, such as salvianolic acid B, tanshinone IIA, and cryptotanshinone, protect vascular endothelial cells by reducing oxidative stress and ameliorating cellular damage [14,15,16]. Additionally, they enhance anticoagulant and fibrinolytic activity, inhibit platelet activation and coagulation, and promote vasodilation [17]. The active ingredients in Hawthorn, including quercetin, chrysin, and hawthorn polysaccharides, can regulate bile acid metabolism [18], ameliorate inflammation and oxidative stress [19], and improve intestinal flora dysbiosis and hepatic metabolic disorders in mice [20]. Collectively, existing evidence substantiates the therapeutic potential of the Danshen & Hawthorn herb pair in the treatment of NAFLD.
In this study, the TCMSP and TCMIP databases were utilized to screen the active components and related targets of the “Danshen & Hawthorn” herb pair. The top ten components ranked by Degree value (a network metric reflecting the number of direct connections a node has to others, where a higher value indicates greater importance [21]) were identified as key active components. Subsequently, potential targets associated with NAFLD were retrieved from the OMIM and GeneCards databases. The intersection between these disease targets and the component-related targets was visualized using a Venn diagram, revealing 27 overlapping genes. These intersecting targets were prioritized based on confidence scores in the STRING database, and the top ten were selected as core targets for further analysis. GO and KEGG enrichment analyses were performed to elucidate the potential mechanisms. The GO analysis indicated that the treatment of NAFLD involves biological processes such as bio-regulation, metabolic regulation, and signaling pathways; cellular components including cellular anatomical entities and protein-containing complexes; and molecular functions like molecular conduction activity, catalytic activity, and ATP-dependent activity. KEGG pathway analysis revealed significant enrichment in pathways related to cancer, NAFLD, the IL-17 signaling pathway, the TNF signaling pathway, the NF-κB signaling pathway, lipid and atherosclerosis, the AMPK signaling pathway, and the PPAR signaling pathway. The IL-17 signaling pathway is implicated as IL-17, a pro-inflammatory cytokine primarily secreted by activated CD4+ T cells, can induce the release of inflammatory factors from epithelial and endothelial cells, thereby promoting inflammatory responses and tissue damage [22]. The TNF signaling pathway primarily regulates the expression of inflammatory genes, including cytokines, chemokines, and adhesion factors, through its downstream effects. TNF can activate the NF-κB pathway [23]. NF-κB, named for its ability to bind the κB enhancer element in the immunoglobulin κ light-chain gene, responds to diverse extracellular signals. Although the canonical NF-κB pathway is rapidly activated, the ensuing transcriptional response is often transient; this pathway primarily regulates pro-inflammatory genes and constitutes a core signaling axis mediating inflammation [24]. The PPAR pathway plays a crucial role in lipid metabolism and inflammatory responses. Its upstream regulators include various endogenous lipid metabolites, inflammatory factors, and insulin [25]. Furthermore, the transcription factor NRF2 can activate downstream antioxidant genes by upregulating PPARγ expression [26]. Activation of the PPAR pathway enhances reverse cholesterol transport and reduces lipid accumulation, conferring cellular protection [27]. The AMPK pathway is a central regulator of cellular energy homeostasis [28]. AMPK responds to low ATP levels and, upon activation, promotes ATP-generating processes such as fatty acid oxidation [29] and autophagy [30]. Conversely, it inhibits ATP-consuming biosynthetic processes including gluconeogenesis, lipid synthesis, and protein synthesis [31]. AMPK exerts these effects either by directly phosphorylating metabolic enzymes or by transcriptionally regulating metabolism through the phosphorylation of transcription factors, co-activators, and co-repressors [32]. In summary, integrated with literature evidence, our analysis suggests that the therapeutic effects of “Danshen & Hawthorn” on NAFLD may involve the IL-17, TNF, NF-κB, AMPK, and PPAR signaling pathways, which are collectively associated with cytoprotection, attenuation of inflammatory responses, reduction of lipid accumulation, and promotion of lipid metabolism. However, the intricate interactions within this network warrant further experimental validation.
To investigate the interactions between the key components and core targets, molecular docking was performed. The top five component-target pairs, ranked by their LibDock scores, were selected for docking visualization. Subsequently, the top three pairs underwent molecular dynamics (MD) simulations and alanine scanning to identify critical binding residues. The top five pairs were PPARα-Quercetin, PPARα-Eburicoic Acid, HMOX1-Dehydroeburicoic Acid, HMOX1-Eburicoic Acid, and PPARγ-Quercetin. Among these, the complexes of PPARα-Quercetin, PPARα-Eburicoic Acid, and HMOX1-Dehydroeburicoic Acid were subjected to 100 ns MD simulations. The results demonstrated that the root-mean-square deviation (RMSD) values for all three complexes stabilized within 3 Å, confirming the stability of the docking poses and reliability of the molecular docking results. The root-mean-square fluctuation (RMSF) analysis further provided insights into the flexibility of protein residues upon ligand binding. Alanine scanning was then performed on these three complexes [33]. This technique assesses the contribution of specific residues to protein-ligand interactions by mutating them to alanine and evaluating the resulting change in binding energy, thereby identifying residues critical for the interaction. The integrated analysis from molecular docking, protein-ligand interaction examination, and alanine scanning revealed the following potential key residues: For PPARα-Quercetin: PHE 207, TYR 134, HIS 25, GLY 143, and SER 53. For PPARα-Eburicoic Acid: ILE 317, MET 355, and PHE 273. For HMOX1-Dehydroeburicoic Acid: ILE 354, PHE 351, TYR 314, SER 280, and HIS 440. Concurrently, the efficacy of the Danshen & Hawthorn extract was evaluated using in vitro (cell-based) and in vivo (zebrafish) models. The extract significantly ameliorated high-fat diet-induced lipid accumulation in both models. The mRNA expression levels of PPARα, TNF-α, and IL-17 were quantified by qPCR. High-fat induction significantly reduced PPARα expression in both models. Given that PPARα is a master regulator of hepatic lipid metabolism and metabolic stress [34], controlling lipid homeostasis by regulating lipid synthesis and catabolism, its downregulation indicated disrupted lipid homeostasis. Treatment with the Danshen & Hawthorn extract reversed this decrease in PPARα, suggesting its intervention in lipid accumulation via PPARα activation. TNF-α, a pivotal cytokine in the pathogenesis of NAFLD and associated steatohepatitis and fibrosis, was significantly upregulated in both high-fat models compared to the blank controls, indicating enhanced inflammation. Existing studies confirm that elevated TNF-α levels are associated with the progression of NAFLD [35]. Treatment with Danshen & Hawthorn reduced TNF-α mRNA levels, demonstrating its anti-inflammatory effect via suppression of TNF-α expression. Similarly, IL-17, a classical inflammatory cytokine, was elevated in the high-fat models. Previous studies have linked obesity to increased activation of the IL-17 axis, which can exacerbate liver injury, confirming the role of IL-17 in NAFLD [36]. Administration of Danshen & Hawthorn reduced IL-17 mRNA expression, indicating its ability to mitigate inflammation in NAFLD by downregulating IL-17.
There are some limitations in this study, because we did not include the setting of positive drug in the model, so we cannot understand the comparison of the efficacy between Danshen and negative drug. However, Danshen and negative drug (Atorvastatin) showed similar effects in alleviating TC and TG [37]. In addition, in the mechanism validation, we only verified the changes in the genes related to NAFLD, without further validation such as gene knockout or overexpression model. At the same time, zebrafish was used for preliminary verification in vivo, and no mice were selected for systematic in vivo verification. Future studies should conduct more systematic in vivo mechanistic studies, as well as “one-to-one” or “one-to-many” binding studies on specific drug components and proteins to clarify their specific contributions and reveal the overall therapeutic mechanism of Danshen and Hawkthorn in NAFLD.
In summary, this study integrated network pharmacology, molecular docking, MD simulations, and alanine scanning to identify feasible key residues within relevant pathways and targets for “Danshen & Hawthorn” in treating NAFLD, thereby refining the potential mechanism of action. The in vitro and in vivo experiments collectively demonstrated that Danshen & Hawthorn intervention in NAFLD is associated with the amelioration of lipid accumulation and reduction of inflammation.
Author Contributions
Z.Z.: Writing—original draft, Visualization, Software, Methodology, Investigation. W.T.: Writing—review & editing. Q.J.: Methodology, Data curation. Y.L.: Methodology, Data curation. Y.Y.: Methodology, Data curation. H.L.: Methodology, Data curation. J.Y.: Writing—review & editing, Supervision, Conceptualization. S.W.: Writing—review & editing, Supervision, Conceptualization, Funding acquisition. All authors have read and agreed to the published version of the manuscript.
Funding
This study was supported by the Natural Science Foundation of Shandong Province under Grant (ZR2024QH063).
Institutional Review Board Statement
Not applicable
Informed Consent Statement
Not applicable.
Data Availability Statement
The authors will supply the relevant data in response to reasonable requests.
Conflicts of Interest
The authors declare no conflict of interest.
Use of AI and AI-Assisted Technologies
No AI tools were utilized for this paper.
References
- 1.
Friedman, S.L.; Neuschwander-Tetri, B.A.; Rinella, M.; et al. Mechanisms of NAFLD development and therapeutic strategies. Nat. Med. 2018, 24, 908–922. https://doi.org/10.1038/s41591-018-0104-9.
- 2.
Powell, E.E.; Wong, V.W.; Rinella, M. Non-alcoholic fatty liver disease. Lancet 2021, 397, 2212–2224. https://doi.org/10.1016/S0140-6736(20)32511-3.
- 3.
Souto, K.P.; Meinhardt, N.G.; Ramos, M.J.; et al. Nonalcoholic fatty liver disease in patients with different baseline glucose status undergoing bariatric surgery: Analysis of intraoperative liver biopsies and literature review. Surg. Obes. Relat. Dis. Off. J. Am. Soc. Bariatr. Surg. 2018, 14, 66–73. https://doi.org/10.1016/j.soard.2017.09.527.
- 4.
Chen, H.; Zhou, Y.; Hao, H.; et al. Emerging mechanisms of non-alcoholic steatohepatitis and novel drug therapies. Chin. J. Nat. Med. 2024, 22, 724–745. https://doi.org/10.1016/S1875-5364(24)60690-4.
- 5.
Wei, B.; Sun, C.; Wan, H.; et al. Bioactive components and molecular mechanisms of Salvia miltiorrhiza Bunge in promoting blood circulation to remove blood stasis. J. Ethnopharmacol. 2023, 317, 116697. https://doi.org/10.1016/j.jep.2023.116697.
- 6.
Cicero, A.F.G.; Colletti, A.; Bellentani, S. Nutraceutical Approach to Non-Alcoholic Fatty Liver Disease (NAFLD): The Available Clinical Evidence. Nutrients 2018, 10, 1153. https://doi.org/10.3390/nu10091153.
- 7.
Zheng, L.; Li, B.; Yuan, A.; et al. TFEB activator tanshinone IIA and derivatives derived from Salvia miltiorrhiza Bge. Attenuate hepatic steatosis and insulin resistance. J. Ethnopharmacol. 2024, 335, 118662. https://doi.org/10.1016/j.jep.2024.118662.
- 8.
Zhu, J.; Guo, J.; Liu, Z.; et al. Salvianolic acid A attenuates non-alcoholic fatty liver disease by regulating the AMPK-IGFBP1 pathway. Chem. Biol. Interact. 2024, 400, 111162. https://doi.org/10.1016/j.cbi.2024.111162.
- 9.
Hong, M.; Li, S.; Wang, N.; et al. A Biomedical Investigation of the Hepatoprotective Effect of Radix salviae miltiorrhizae and Network Pharmacology-Based Prediction of the Active Compounds and Molecular Targets. Int. J. Mol. Sci. 2017, 18, 620. https://doi.org/10.3390/ijms18030620.
- 10.
Lu, M.; Zhang, L.; Pan, J.; et al. Advances in the study of the vascular protective effects and molecular mechanisms of hawthorn (Crataegus anamesa Sarg.) extracts in cardiovascular diseases. Food Funct. 2023, 14, 5870–5890. https://doi.org/10.1039/d3fo01688a.
- 11.
Li, T.; Wang, H.; Dong, S.; et al. Protective effects of maslinic acid on high fat diet-induced liver injury in mice. Life Sci. 2022, 301, 120634. https://doi.org/10.1016/j.lfs.2022.120634.
- 12.
Wang, T.; Wang, D.; Ding, Y.; et al. Targeting Non-Alcoholic Fatty Liver Disease with Hawthorn Ethanol Extract (HEE): A Comprehensive Examination of Hepatic Lipid Reduction and Gut Microbiota Modulation. Nutrients 2024, 16, 1335. https://doi.org/10.3390/nu16091335.
- 13.
Hamza, A.A.; Lashin, F.M.; Gamel, M.; et al. Hawthorn Herbal Preparation from Crataegus oxyacantha Attenuates In Vivo Carbon Tetrachloride—Induced Hepatic Fibrosis via Modulating Oxidative Stress and Inflammation. Antioxidants 2020, 9, 1173. https://doi.org/10.3390/antiox9121173.
- 14.
Liu, Q.; Shi, X.; Tang, L.; et al. Salvianolic acid B attenuates experimental pulmonary inflammation by protecting endothelial cells against oxidative stress injury. Eur. J. Pharmacol. 2018, 840, 9–19. https://doi.org/10.1016/j.ejphar.2018.09.030.
- 15.
Zhou, Z.Y.; Shi, W.T.; Zhang, J.; et al. Sodium tanshinone IIA sulfonate protects against hyperhomocysteine-induced vascular endothelial injury via activation of NNMT/SIRT1-mediated NRF2/HO-1 and AKT/MAPKs signaling in human umbilical vascular endothelial cells. Biomed. Pharmacother. 2023, 158, 114137. https://doi.org/10.1016/j.biopha.2022.114137.
- 16.
Cui, W.B.; Zhang, Z.P.; Bai, X.; et al. Cryptotanshinone Alleviates Oxidative Stress and Reduces the Level of Abnormally Aggregated Protein in Caenorhabditis elegans AD Models. Int. J. Mol. Sci. 2022, 23, 10030. https://doi.org/10.3390/ijms231710030.
- 17.
Zhang, T.; Liu, M.; Gao, Y.; et al. Salvianolic acid B inhalation solution enhances antifibrotic and anticoagulant effects in a rat model of pulmonary fibrosis. Biomed. Pharmacother. 2021, 138, 111475. https://doi.org/10.1016/j.biopha.2021.111475.
- 18.
Wang, S.; Sheng, F.; Zou, L.; et al. Hyperoside attenuates non-alcoholic fatty liver disease in rats via cholesterol metabolism and bile acid metabolism. J. Adv. Res. 2021, 34, 109–122. https://doi.org/10.1016/j.jare.2021.06.001.
- 19.
Yang, H.; Yang, T.; Heng, C.; et al. Quercetin improves nonalcoholic fatty liver by ameliorating inflammation, oxidative stress, and lipid metabolism in db/db mice. Phytother. Res. 2019, 33, 3140–3152. https://doi.org/10.1002/ptr.6486.
- 20.
Zhou, Y.; Wang, M.; Wang, Z.; et al. Polysaccharides from hawthorn fruit alleviate high-fat diet-induced NAFLD in mice by improving gut microbiota dysbiosis and hepatic metabolic disorder. Phytomed. Int. J. Phytother. Phytopharm. 2025, 139, 156458. https://doi.org/10.1016/j.phymed.2025.156458.
- 21.
Li, X.; Liu, Z.; Liao, J.; et al. Network pharmacology approaches for research of Traditional Chinese Medicines. Chin. J. Nat. Med. 2023, 21, 323–332. https://doi.org/10.1016/S1875-5364(23)60429-7.
- 22.
Kumar, P.; Monin, L.; Castillo, P.; et al. Intestinal Interleukin-17 Receptor Signaling Mediates Reciprocal Control of the Gut Microbiota and Autoimmune Inflammation. Immunity 2016, 44, 659–671. https://doi.org/10.1016/j.immuni.2016.02.007.
- 23.
Low, J.T.; Christie, M.; Ernst, M.; et al. Loss of NFKB1 Results in Expression of Tumor Necrosis Factor and Activation of Signal Transducer and Activator of Transcription 1 to Promote Gastric Tumorigenesis in Mice. Gastroenterology 2020, 159, 1444–1458.e15. https://doi.org/10.1053/j.gastro.2020.06.039.
- 24.
Wu, N.; McDaniel, K.; Zhou, T.; et al. Knockout of microRNA-21 attenuates alcoholic hepatitis through the VHL/NF-κB signaling pathway in hepatic stellate cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2018, 315, G385–G398. https://doi.org/10.1152/ajpgi.00111.2018.
- 25.
Paumelle, R.; Haas, J.T.; Hennuyer, N.; et al. Hepatic PPARα is critical in the metabolic adaptation to sepsis. J. Hepatol. 2019, 70, 963–973. https://doi.org/10.1016/j.jhep.2018.12.037.
- 26.
Shou, J.W.; Li, X.X.; Tang, Y.S.; et al. Novel mechanistic insight on the neuroprotective effect of berberine: The role of PPARδ for antioxidant action. Free Radic. Biol. Med. 2022, 181, 62–71. https://doi.org/10.1016/j.freeradbiomed.2022.01.022.
- 27.
Zhao, J.; Sun, Y.; Yuan, C.; et al. Quercetin ameliorates hepatic fat accumulation in high-fat diet-induced obese mice via PPARs. Food Funct. 2023, 14, 1674–1684. https://doi.org/10.1039/d2fo03013f.
- 28.
Ke, R.; Xu, Q.; Li, C.; et al. Mechanisms of AMPK in the maintenance of ATP balance during energy metabolism. Cell Biol. Int. 2018, 42, 384–392. https://doi.org/10.1002/cbin.10915.
- 29.
Lopez-Mejia, I.C.; Lagarrigue, S.; Giralt, A.; et al. CDK4 Phosphorylates AMPKα2 to Inhibit Its Activity and Repress Fatty Acid Oxidation. Mol. Cell 2024, 84, 2397. https://doi.org/10.1016/j.molcel.2024.05.021.
- 30.
Wang, B.; Shi, Y.; Chen, J.; et al. High glucose suppresses autophagy through the AMPK pathway while it induces autophagy via oxidative stress in chondrocytes. Cell Death Dis. 2021, 12, 506. https://doi.org/10.1038/s41419-021-03791-9.
- 31.
Qi, R.; Wang, J.; Wang, Q.; et al. MicroRNA-425 controls lipogenesis and lipolysis in adipocytes. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2019, 1864, 744–755. https://doi.org/10.1016/j.bbalip.2019.02.007.
- 32.
Grandjean, G.; de Jong, P.R.; James, B.; et al. Definition of a Novel Feed-Forward Mechanism for Glycolysis-HIF1α Signaling in Hypoxic Tumors Highlights Aldolase A as a Therapeutic Target. Cancer Res. 2016, 76, 4259–4269. https://doi.org/10.1158/0008-5472.CAN-16-0401.
- 33.
Clackson, T.; Wells, J.A. A hot spot of binding energy in a hormone-receptor interface. Science 1995, 267, 383–386. https://doi.org/10.1126/science.7529940.
- 34.
Kim, D.; Ha, S.K.; Gonzalez, F.J. CBFA2T3 Is PPARA Sensitive and Attenuates Fasting-Induced Lipid Accumulation in Mouse Liver. Cells 2024, 13, 831. https://doi.org/10.3390/cells13100831.
- 35.
Vachliotis, I.D.; Polyzos, S.A. The Role of Tumor Necrosis Factor-Alpha in the Pathogenesis and Treatment of Nonalcoholic Fatty Liver Disease. Curr. Obes. Rep. 2023, 12, 191–206. https://doi.org/10.1007/s13679-023-00519-y.
- 36.
Harley, I.T.; Stankiewicz, T.E.; Giles, D.A.; et al. IL-17 signaling accelerates the progression of nonalcoholic fatty liver disease in mice. Hepatology 2014, 59, 1830–1839. https://doi.org/10.1002/hep.26746.
- 37.
Zhang, X.; Zhang, J.; Zhou, Z.; et al. Integrated network pharmacology, metabolomics, and transcriptomics of Huanglian-Hongqu herb pair in non-alcoholic fatty liver disease. J. Ethnopharmacol. 2024, 325, 117828. https://doi.org/10.1016/j.jep.2024.117828.

This work is licensed under a Creative Commons Attribution 4.0 International License.


